Research Article - Der Pharma Chemica ( 2018) Volume 10, Issue 5
Evaluation of Antioxidant Property and Binding Constant with DNA using Quinones by Electrochemical Technique
Ramaraj R Deepa1, Abraham A Daniel Arulraj2, Abdul Kather A. Sheik Mideen3, Rathinasamy R Gandhidasan4 and Vairathevar Sivasamy VS Vasantha1*
1Department of Natural product chemistry, Madurai Kamaraj University, Madurai- 625 021, India
2National Postac Fellow, Department of chemistry, SRM University, Kattankulathur, Kancheepuram District, Chennai-603203, India
3Department of Chemistry, Sathyabama University, Jeppiaar Nagar,Chennai-600119,India
4Department of chemistry, K. L. N. College of Engineering, Pottapalayam, Sivagangai District, Madurai-630612, India
- *Corresponding Author:
- Vairathevar Sivasamy VS Vasantha
Department of Natural product chemistry
Madurai Kamaraj University
Madurai- 625 021, India
Abstract
Antioxidant property and interaction with Deoxyribonucleic Acid (DNA) of the following four quinones: Mansanone-D, Mansanone-H, Sissoidenone and 4-methoxy dalbergione, were investigated at pH=7.4 using cyclic voltammetry technique. The parameters like binding constant, binding site size and binding free energy were also determined from voltammetric data. On comparison of binding constant and binding site size among all quinones, Mansanone-H reveals strong binding capacity (1.04 x 109) with DNA than others. The negative sign of binding constants (ΔG) indicate the spontaneity of binding of quinones with DNA. The electrochemical behavior of DPPH radical was investigated in Dimethyl Sulfoxide (DMSO) and used this radical to determine antioxidant activity of quinones by cyclic voltammetry method. The scavenging effect of the radical was monitored by the decrease in peak currents with increase in the concentration of quinones. The binding constant and Gibbs free energy values were calculated from voltammetric data.
Graphical Abstract
Keywords
Isolated quinones, DPPH, Calf thymus DNA, Cyclic voltammetry, Binding constant
Introduction
Plant cells produce two types of metabolites, primary metabolites (carbohydrate, proteins and lipids) and secondary metabolites (Alkaloids, phenolics, stereols, steroids, essentialoils, lignins, tannins, etc.). The secondary metabolites act as defense chemicals. Many thousands of secondary metabolites have been isolated from plants and many of them have powerful physiological effects in humans. They contain a wide variety of free radical scavenging molecules, such as flavonoids, anthocyanins, carotenoids, dietary glutathionine, vitamins, endogenous metabolites, terpenoids, phenolic acids, lignins, stilbenes, tannins, flavonoids, Quinones, coumarins, alkaloids, amines, betalains, etc., which are rich in antioxidant activity [1,2].
Reactive Oxygen species is a collective term used for oxygen derived free radicals such as super oxide (O2●−), hydroxyl (OH●), hydrogen peroxide (H2O2), nitric oxide and peroxyl (ROO●) radicals [3]. Free radical is a chemical species which contains an unpaired electron spinning on the peripheral layer around the nucleus. Free radicals generated from the oxygen are called Reactive Oxygen Species (ROS) which cause damage to other molecules by extracting electrons from them in order to attain stability. Most of the free radicals are very unstable and thus highly reactive in nature. Nowadays, it is reported that a reactive oxygen species (ROS) is implicated in a wide range of human diseases such as atherosclerosis and certain cancers.
For in vitro characterization, several methods have been proposed for the detection of antioxidants such as photometric, fluorimetric, chromatographic and electrochemical methods.
By using different scavenging reagents like DPPH, ABTS, FRAP, PFRAP, CUPRAC, ORAC, HORAC and TRAP, the antioxidant activity of natural products were studied by using different spectroscopic methods. Among them, the DPPH is a rapid, simple, easy, inexpensive and widely used method to measure the ability of compounds to act as antioxidant.
DPPH molecules are utilized in aqueous and nonpolar organic solvents and can be used to examine both hydrophilic and lipophilic antioxidants. The results are highly reproducible and comparable to other free radical scavenging methods. Recently, the electrochemical methods such as cyclic voltammetry, amperometry, biamperometry techniques etc., have been used to determine the antioxidant activities of plant materials [4]. The main advantages of electrochemical methods are rapid, simple and sensitive alternative method in the analysis of bioactive compounds associated with the scavenging of the radicals as well as the antioxidant capacity itself. They are low-cost and usually do not require time consuming sample preparation. Significant work has been done on estimating antioxidant activity of different substrates, limited literature is available on the electrochemistry of 2,2-diphenyl-1-picrylhydrazyl radical (DPPH). It is of interest to note that in quinone type compound, no electrochemical work is found which uses DPPH as target radical to explore antioxidant activity.
Protection of DNA from being damaged has become an important issue in both chemistry and medicinal field. DNA damage caused by oxygen‐derived species including free radicals is the most frequent type encountered by aerobic cells which is called oxidative DNA damage. Further, oxidative DNA damage plays an important role in a number of disease processes such as carcinogenesis, cardiovascular, neurodegenerative and inflammatory diseases [5]. Therefore several techniques have been employed to study the binding of small molecules to DNA including, viscometry, UV-Vis spectroscopy [6], isothermal calorimetric titration, luminescence, electrophoresis [7], fluorescence [8] and electroanalytical method. Among various methods, electrochemical method is widely used in drug–DNA binding studies due to their advantages like high sensitivity, efficient selectivity, cost affectivity, more reliability, extensive versatility and fast detection ability. There are mainly three possible binding modes between DNA and small molecules: (i) intercalative binding that small molecules intercalate into the base pairs of nucleic acids [9] (ii) groove binding in which the small molecules bound on nucleic acids are located in the major or minor groove and (iii) longrange assembly on the molecular surfaces of nucleic acids so that the small molecules are not related to the groove structure of the nucleic acids [10]. The intercalative binding is stronger than other two binding modes because the surface of intercalative molecule is sandwiched between the aromatic and heterocyclic base pairs of DNA [11,12].
The recent years, growing interest has arisen in electrochemical investigations of interactions between nucleic-acid-binding molecules and DNA using mercury or carbon especially electrodes [13-16]. Bard et al. firstly applied electrochemical methods to investigate the interaction between DNA and the electrochemical active metal complexes in solution [17]. These investigations are mainly based on the differences in the electrochemical behaviors of the nucleic acid-binding molecules in the absence and presence of double-stranded DNA including the shifts of the formal potentials of the redox couple caused by the intercalation of nucleic-acid-binding molecules into DNA double helix and the reduction of peak current resulting from the dramatic decrease in the apparent diffusion coefficient of the nucleic-acid-binding molecule after association with double-stranded DNA [18].
In traditional medicine all over the world, plants which are rich in quinones are used for the treatment of a variety of diseases. Quinone constitutes one of the well-known groups of naturally occurring organic compounds and structurally diverse class of phenolic compounds with a wide range of pharmacological properties, which are the basis for different applications in the broad field of pharmacy and medicine. Major attraction of the researchers is that many of the drugs clinically approved or still in clinical trials against cancer are quinone related compounds. Quinones, present in several living organisms, participate to various biological processes. Quinones can also interact with DNA, which has been demonstrated in many works. Some quinines can interact with nucleobases from DNA and inhibit replication [19] or bind to double-stranded DNA (dsDNA) by intercalation [20,21] or groove binding [22]. Since the concept of intercalation into DNA was first formulated by Lerman [23] in 1961. These properties make quinoid compounds attractive for anti-cancer drugs [24,25].
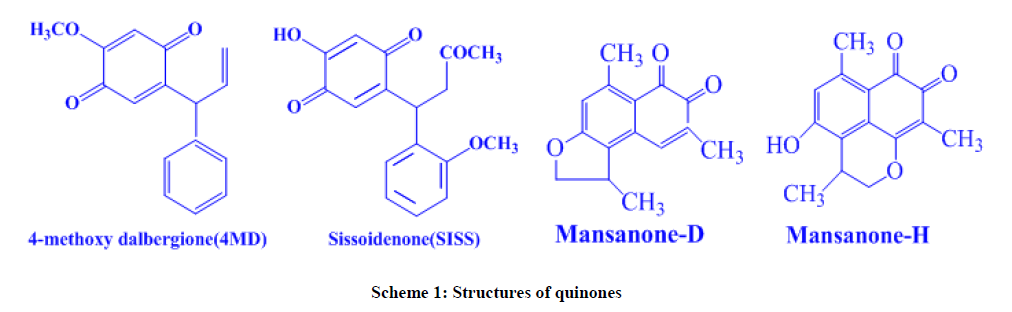
The present work is the result of an attempt to develop a simple electrochemical method (cyclic voltammetry) for determining antioxidant activity and interactions between DNA of four naturally occurring quinones (Scheme 1) namely 4-methoxydalbergione(4MD) [26], Sissoidenone (Siss) [26], Mansanone-D (MD) [27] and Mansanone-H(MH) [27]. The binding constants values were calculated for above molecules and compared with earlier reports. The value of the binding constant of MH-DNA is found to be very high compare to other three and compared with earlier reports.
Materials and Methods
Cyclic voltammetric measurements were carried out using CH analyzer. An electrochemical cell with conventional three electrode system was employed in all electrochemical measurements. Glassy carbon (3 mm dia) and platinum foil electrodes were used as working and counter electrodes, respectively. A saturated silver–silver chloride electrode was used as reference electrode. Before each experiment, the surface of GCE was polished with alumina powder followed by thorough rinsing with distilled water. All solutions were purged with pure nitrogen gas for 10 min and an inert atmosphere was maintained during the experiments.
Dimethylsulphoxide (DMSO) was obtained from Fluka and was used without further purification. Tetrabutyl ammonium perchlorate (TBAP, Fluka (99% purity) was used as supporting electrolyte. 2,2-Diphenyl-1-picrylhydrazyl radical (98%) was purchased from Sigma Aldrich. The natural products such as Mansanone D(MD), and Mansanone H(MH) were isolated from Thepesia populnea, Sissoidenone(Siss), and 4-Methoxy Dalbergione (4MD) were isolated from Dalbergia sissoides. The data corresponding to isolation were already published [26,27]. The highly polymerized double-stranded Calf-thymus DNA sodium salt (7% Na content) was purchased from Sigma Chemical Co. The Tris–HCl (10 mM, pH 7.2) was purchased from sigma Aldrich and dissolved in distilled water. All experiments involving CT-DNA were performed in Tris buffer solution containing 50 mM NaCl/1 mM Tris–HCl at pH 7.2. The concentration of CT-DNA was first determined using UV-Vis spectroscopic technique. From the intensity of absorbance at 260 nm and extinction coefficient value (£260=6600 M-1 cm-1), the concentration of CT-DNA was calculated [28]. The ratio of the absorbance at 260 nm to that at 280 nm is 1.9 (A260/A280> 1.8), which indicates that DNA is sufficiently pure and free from other proteins. For antioxidant activity measurement, stock solutions of 0.1 M TBAP and 0.5 mM DPPH• were prepared in DMSO and Stock solution of DNA was prepared by dissolving the appropriate amounts of DNA in Tris buffer while those of quinones (each of 2.5 mM) were dissolved in DMSO. Both DNA and quinone solutions were stored at 4°C.
Results and Discussion
Electrochemical behavior of quinone
The electrochemical behaviors of sesquiterpenoid orthonaphtho quinones (MD and MH)and para benzoquinones (Siss and 4MD) were studied using cyclic voltammetry technique in 2.5 mM quinones dissolved in 0.1 M TBAP DMSO solutions in the potential range of -1.0 V to +1.0 V at 100 mV/s. Generally in non-aqueous media, quinones undergo reduction by two successive one electron transfer steps in which the first step is reversible, while the second step is quasi reversible [29]. But the results of our experiments indicate that the one quasi reversible redox couple and another irreversible reduction peak are appeared for the compounds 4MD and Siss with peak separation of 143 mV and 170 mV, respectively whereas the cyclic voltammograms of MD and MH show only one reversible redox couple with peak separation of 66 mV and 44 mV, respectively. Similarly the ipa/ipc ratios for 4MD, Siss, MD and MH were calculated as 0.56, 0.55, 1.37 and 0.65, respectively. The parameters derived from cyclic voltammogram of 4MD and Siss indicate obviously one quasi reversible electrochemical step for the formation of semiquinone and another broad cathodic peak corresponds to second electron transfer step for the formation of dianione. But in the case of MD and MH, the peak separation data show that the electrochemical behavior of MD and MH are highly reversible for the formation of dianion. However, the unequal anodic and cathodic peak currents pointed out to the quasi reversible nature of the overall electrochemical process. This may be due to adsorption of oxidized molecules on the electrode surface. Moreover, an additional oxidation peak was observed at higher anodic potential (at +0.7 V) for MH. It resulted from the oxidation of –OH group present at 6th position of MH.
By comparing the ortho and para pair of compounds, the Epc values of ortho-quinone (MD) appears at-0.286 V and the para quinones (Siss) appears at +0.010 V. Within the quinones investigated in the present study, orthoquinones are shown to be more easily oxidized than the para quinones, previous observations of quinones are corroborating with this kind of observation [30]. Very few reports are available for the systematic comparison of ortho and para electrochemical properties [31]. Very recently, Stoppani and Coworkers [32] concluded that based on the calculated LUMO atomic charges and due to electronic asymmetry, the polarity of C=O bond is higher in ortho naphthoquinones than para quinones. This aspect makes the carbonyl carbon atom of ortho compounds more electrons deficient and hence, makes it easier to oxidize than in para quinones.
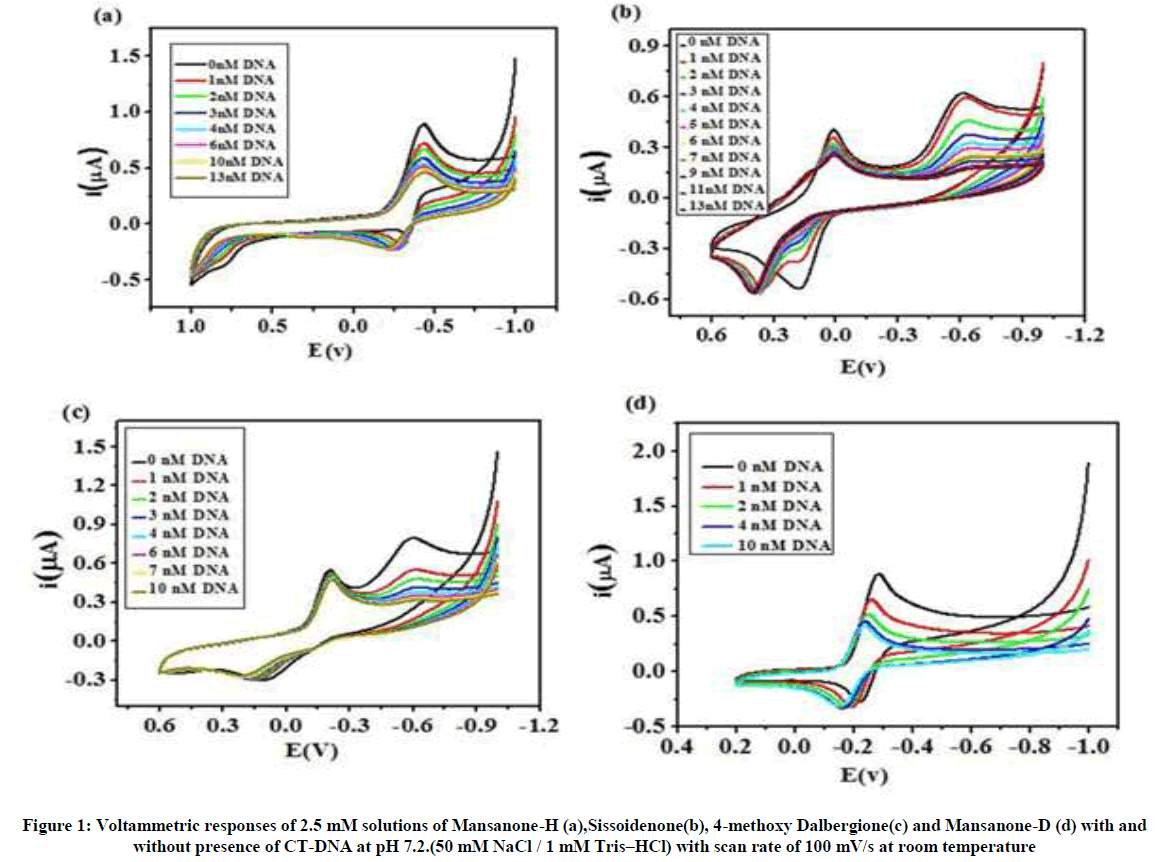
Cyclic Voltammetry investigation of quinones-DNA interaction
In the present study, Cyclic Voltammetry technique was employed to understand the nature of the binding of quinones with CT-DNA. Typical cyclic voltammetric behavior of the above mentioned four quinones, in the absence and presence of CT-DNA, are shown in Figure 1 (a,b,c &d). Figure 1(a) show that the one redox couple (anodic and cathodic peaks are occurred at -385 mV and -429 mV, respectively) and broad peak (0.70 mV) are observed in the cyclic voltammogram for MH. The addition of different concentrations of DNA to the fixed concentration of MH (0.5 mM) caused a decrease in the reduction peak currents with no shift in peak potential. Additionally, the oxidation peak potential shifted (85 mV) to more positive value in the presence of CT-DNA.
As shown in Figure 1(b), two reduction peaks and one oxidation peak at +181 mV and +10 mV and -0.615 V, respectively. When CT-DNA was added into the Siss solution, caused a gradual decrease in the first and second cathodic peak currents with a slight negative shift in the second irreversible cathodic peak potential (from -0.615 V to -0.621 V) whereas the anodic peak potential is shifted to more positive value (from +0.181 V to +0.352 V). As reported previously, the shift of oxidation peak potential in the CV to more positive values, indicating that the intercalation mode of interaction of quinone with dsDNA [33]. Further the shift in second cathodic peak to more negative side can be linked with electrostatic interaction between the anionic phosphate back bone of DNA. Similar kinds of interaction was observed for 9,10-Anthraquinone and 2- Chloromethylanthraquinone [34].
Similarly, in the presence of CT-DNA, two cathodic peak currents of Figure 1(c) are decreased and peak potential of second cathodic wave is shifted to more negative value (from -0.594 V to -0.620 V). The oxidation peak is also shifted to more positive side (30 mV). The results indicating that the interaction of quinone with CT-DNA is electrostatic as well as intercalation modes. So the interactions of compounds Siss and 4MD are expected to alter the DNA replication machinery that may lead to the death of cancerous cells. Figure 1(d) shows that in the presence of CT-DNA caused the decrease in cathodic peak current and anodic peak current left unchanged. However, the reduction and oxidation peak potentials are shifted to more positive value by 25 mV. The behavior could be attributed to perpendicular placement of quinone molecules between the adjacent base pairs of dDNA i. e., intercalation. The decreases in peak currents can be explained in terms of the slow diffusion of the quinone-DNA supramolecular complex due to which the concentration of the free drug which is mainly responsible for the transfer of current, is lowered [35,36].
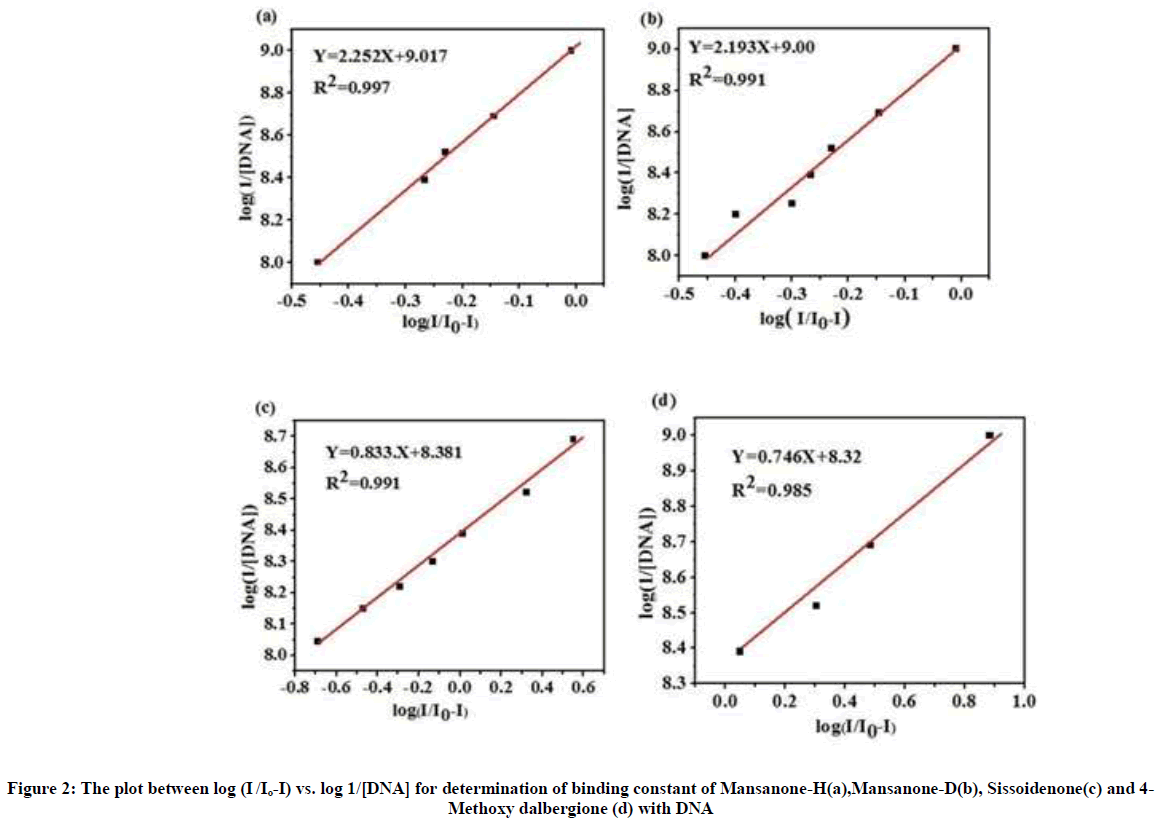
The decay in peak current (Ip) of the quinones by the addition of increasing amount of DNA can be used for the determination of binding constant and binding site size, whereas the shift in peak potential can be used to ascertain the mode of interaction. Based upon the decrease in peak currents MH, MD, Siss and 4MD by the addition of different concentration of DNA, ranging from 1 to 13 nM, the binding constant was calculated from the intercept of the plot of log (1/DNA) versus log (I/Io-I) using the equation 1
log (1/[DNA]) = log K + log (I/(Io-I)) (1)
where, K is the binding constant, Io and I are the peak currents of the drug in the absence and presence of DNA, respectively.
Figure 2 shows the binding constants of MH, MD, Siss and 4MD are calculated as 1.04 x 109, 1.00 x 109, 2.3 x 108 and 2.08 x 108 and respectively. The binding constants values of our all quinones are found to be very higher than the binding constant values of already isolated quinones and flavonoids used as anticancer drugs. For example, the higher binding constant values of Nogalamycin (classical intercalator) is K= 4.44 x 105 [37] and the clinically used anticancer drug Daunorubicin is K=2.44 x 105 [38] which suggests that our compounds can be preferred as better anticancer drug candidature.
From the cyclic voltammetric data acquired from the interaction between quinones and DNA, a simple site binding model was proposed. For the determination of binding site size the following equation was used [39].
Cb/Cf = K ([free base pairs]/s (2)
Whereas is the binding site size in terms of base pairs. Measuring the concentration of DNA in terms of [NP], the concentration of base pairs can be expressed as [DNA]/2. So equation 2 can be re-written as:
Cb/Cf = K([DNA]/2s (3)
Cf and Cb denote the concentrations of free and DNA-bound species, respectively. The ratio of the bound and free drug was determined by the equation given below [40]
Cb/Cf = (I − IDNA)/IDNA (4)
Where IDNA and I, represent the peak currents of the drug with and without DNA.
Putting the value of K as calculated according to above equation 1 the size of binding site of DNA with quinones were calculated and given in Table 1.
| Quinone-DNA Complex | Kb × (M-1) | - ΔG0 (kJmol-1) |
S | R | Mode of binding |
|---|---|---|---|---|---|
| Mansanone H-DNA | 1.04 × 109 | 52.83 | 2.23 | 0.99 | Intercalation |
| Mansanone D-DNA | 1.0 × 109 | 52.73 | 0.69 | 0.99 | Intercalation |
| Sissoidenone-DNA | 2.3 × 108 | 48.99 | 0.23 | 0.99 | Intercalation and electrostatic |
| 4-Methoxy dalbergione -DNA | 2.08 × 108 | 48.73 | 0.32 | 0.98 | Intercalation and electrostatic |
Kb is the binding constant; - ΔG0 is the Gibbs free eneegy values; S and R are the binding site size and correlation coefficient respectively
Table 1: Binding constant, change in standard Gibbs free energy of reaction, binding site size and correlation coefficient of quinone‐DNA complexes calculate from cyclic voltammetry data
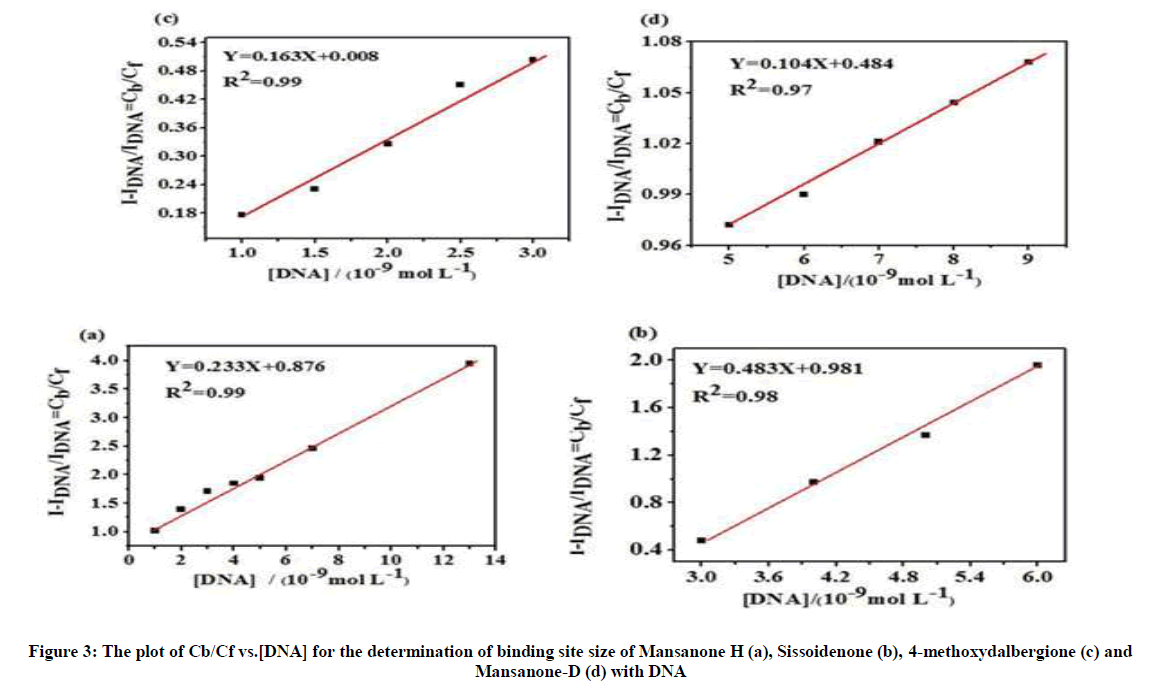
Figure 3 depicts the plot of Cb/Cf vs. [DNA] for determination of binding site size of MH (a), Siss (b), 4MD (c) and MD (d) with DNA at 37°C.
The binding site size of MH, Siss, 4MD and MD are calculated as 2.23 bp, 0.23 bp, 0.32 bp and 0.69 bp, respectively. The binding site size values of Siss, 4MD and MD with CT-DNA are low compare to binding site size value of MH. The binding site number implies only a binding site per two base pairs, giving evidence for an intercalation of MH into DNA according to the neighbor-exclusion model, since groove binding and electrostatic binding usually results in significantly higher binding site number [41,42]. The binding site size determined shows that MH (n=2.23) covers more than two base pairs upon binding with DNA.
Electrochemical behavior of DPPH radical in DMSO
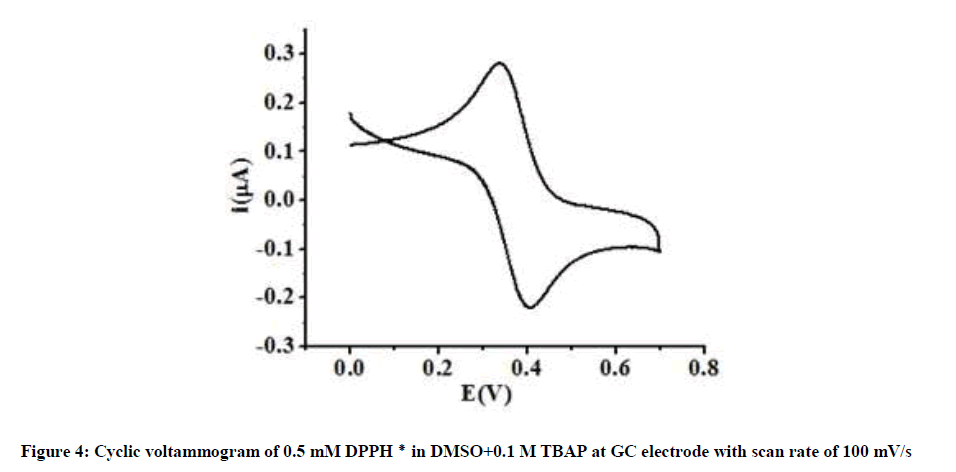
To study the electrochemical behavior of DPPH radical, 0.5 mM solution of DPPH radical was prepared by dissolving in DMSO + 0.1 M TBAP solution and cyclic voltammogram was recorded in the potential range of 0.0 V to + 0.7 V at scan rate of 100 mV/s. The cyclic voltammogram exhibits a single reversible redox couple with formal potential at -0.406 V. Here, the DPPH radical undergoes one electron reduction to form the anion DPPH ¯ and one electron oxidation to form the cation DPPH+. This voltammetric response is agreement with earlier reports [43-45]. The average peak separation (ΔEp) is 66 mV and the peak current ratio is 1.00 (Figure 4).
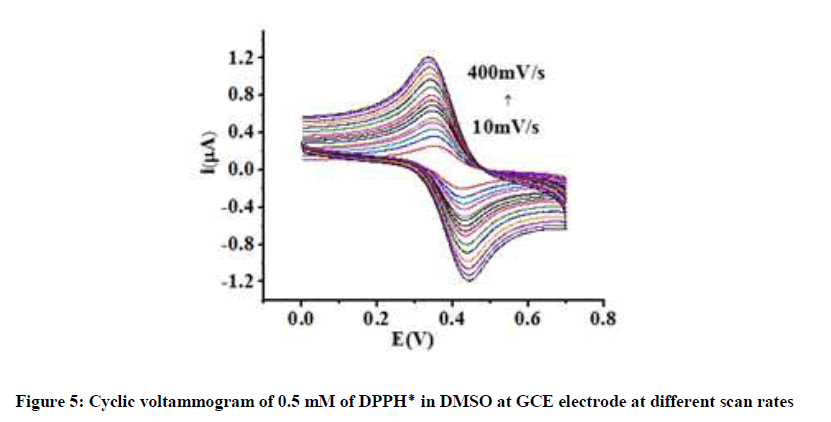
These values indicate that electrochemical behavior of DPPH radical in DMSO is highly reversible. It is further confirmed by the constant values of peak potential observed on varying the scan rate (Figure 5).
Voltammetric behavior of DPPH radical in the presence of quinones
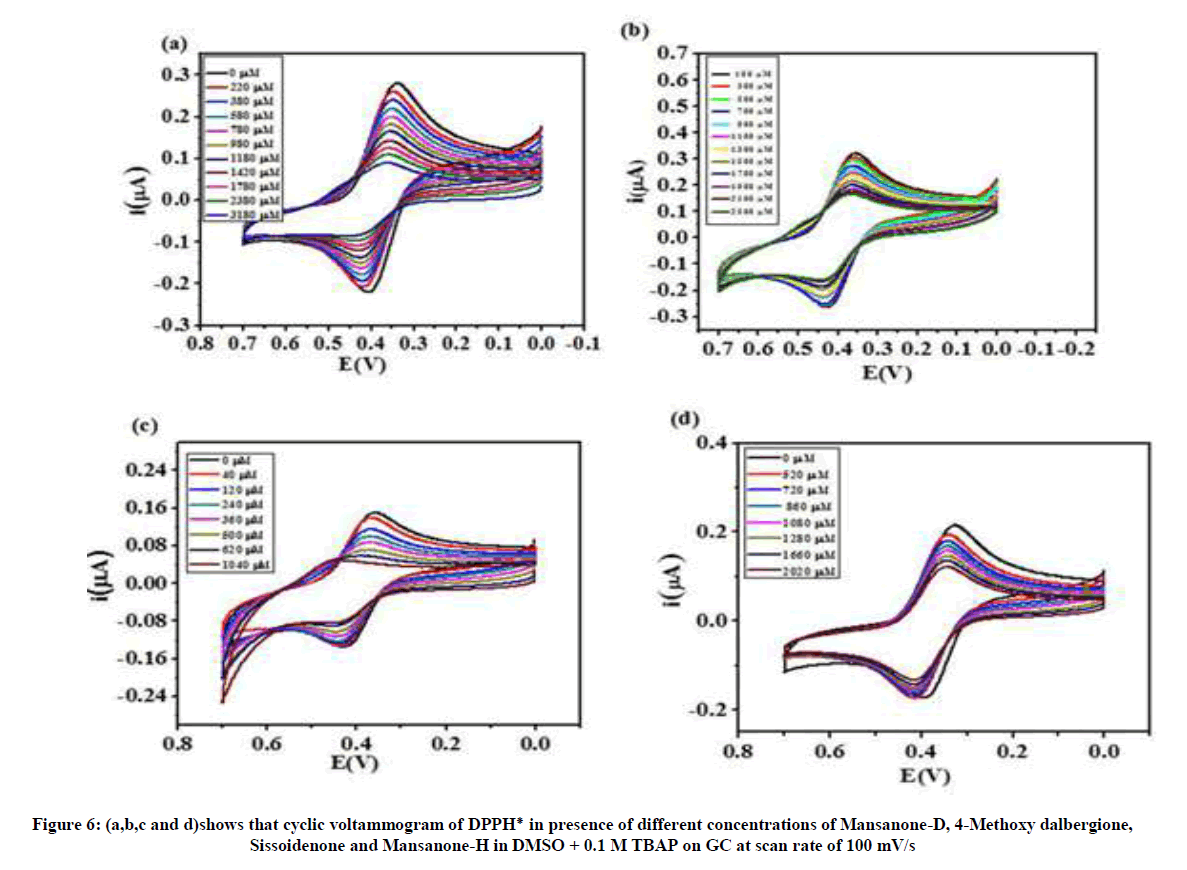
A cyclic voltammogram of DPPH radical was recorded in DMSO at pH=9.5 in 0.1 M TBAP and in the potential range of + 0 V to + 0.7 V at a scan rate of 100 mV/S. The quinones were added in incremental amounts in the concentration range of 20 μM to 3500 μM and the CV response was recorded for each addition. The antioxidant activity of the quinones was estimated from the anodic current changes observed in the voltammograms. Results for four compounds are presented in Figure 6.
Figure 6 shows on adding a quinone to a solution containing DPPH٭ causes a decreases the anodic and cathodic peak currents and the anodic peak potential of the compound (MD, Siss, MH) is slightly shifted towards positive side which is due to the scavenging activity of the added quinone.
The decrease in the anodic current is attributed to the decrease of the radical concentration and decrease in the cathodic current indicates electron transfer mechanism from the antioxidant to the DPPH radical immediately followed by a proton transfer. The 4MD show no shift in anodic peak potential, only decrease in both anodic and cathodic peak currents. The increase of the antioxidant substrate (quinone) concentration leads to a decrease DPPH٭ anodic peak current. Afterwards, this decrease in the current gets gradually smaller and becomes constant at certain concentration value (e. g. 3180 μM for MD, 2260 μM for MH, 2500 μM for 4MD and 1040 μM for siss). This current decrease reflects the scavenging of the quinone and it provides a direct measure of antioxidant capacity [46]. Some of the previous cited authors established an interesting relationship between electrochemical behavior of the antioxidant compounds and their resultant ‘‘antioxidant power”, where ‘‘low oxidation potential” corresponds to ‘‘high antioxidant power”. Recently some researchers have already reported a direct correlation between antioxidant activity of flavonoids and their first oxidation peak potentials [47,48]. The oxidation peak potentials of MD, MH, Siss and 4MD are - 0.220 V, -0.385 V, +0.181 V and +0.020 V, respectively.
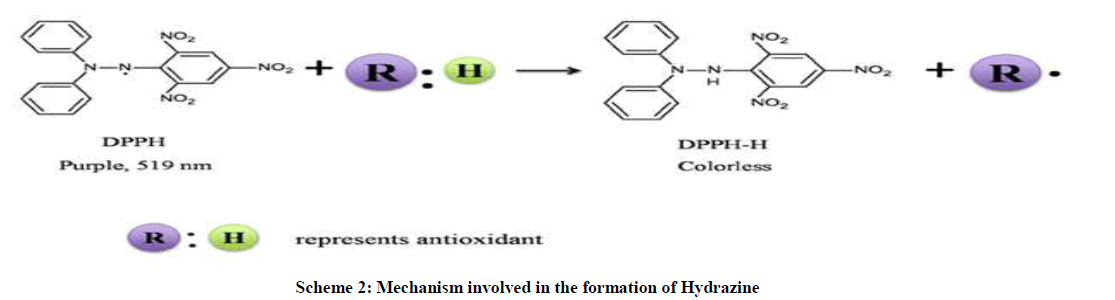
Voltammograms indicate the mechanism of radical-scavenging process. In an electron transfer mechanism DPPH is relatively stable nitrogen centered free radical that easily accepts an electron or hydrogen radical to become a stable diamagnetic molecule. DPPH radicals react with suitable reducing agents as a result of which the electrons become paired off forming the corresponding hydrazine (Scheme 2). The overall mechanism is shown below in equation 5.
DPPH٭+ QH → DPPH - + QH +٭ → DPPH-H+Q٭ (5)
In this mechanism the DPPH٭ extract an electron from the quinone to form mono anion (DPPH-) and second step proton is transferred from DPPH- to form DPPH-H (2,2-diphenyl-1-picrylhydrazine).
Binding constant (Kb)
To quantify the results, the strength of interaction between DPPH radical and antioxidant was estimated in terms of the binding constant Kb. It was calculated using the equation 2.
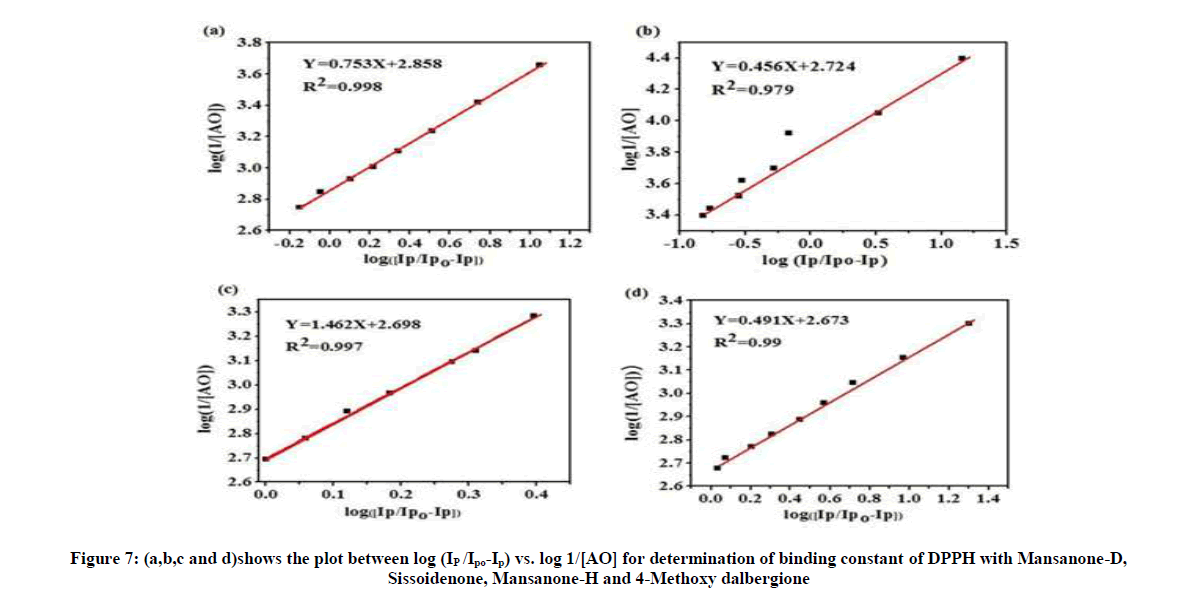
log(1/[AO]) = log Kb + log (Ip/Ipo-Ip) (6)
Where Ipo, Ip =Peak currents of DPPH radical in the absence and presence of additives [AO] = Concentration of antioxidant
Figure 7 shows the plot between log (IP/Ipo-Ip) vs. log1/[AO] for determination of binding constant of DPPH with Mansanone-D, Sissoidenone, Mansanone-H and 4-Methoxy dalbergione.
Thermodynamic parameter (ΔG0) was calculated using the Kb value obtained from equation 7.
ΔG0 = -RT ln Kb (7)
The free energy change (ΔG0) was estimated from equation 7 and the values are listed in Table 2.
| Antioxidant | % decrease (Ipc) | % decrease (Ipa) | Kb x (M-1) | -ΔG0 (kJmol-1) |
|---|---|---|---|---|
| Mansanone-D | 61.59 | 54.12 | 7.21 × 102 | 16.74 |
| Sissoidenone | 46.74 | 22.93 | 5.29 × 102 | 15.95 |
| Mansanone-H | 39.8 | 34.57 | 4.98 × 102 | 15.8 |
| 4-Methoxy dalbergione | 60.02 | 46.39 | 4.70 × 102 | 15.65 |
Ipc and Ipa are the percentage decrease in cathodic and anodic peak current respectively; Kb is the binding constant; -ΔG0 is the standard gibbs free energy
Table 2: Decrease in current values, binding constant and change in standard Gibbs free energy of reaction (ΔG0) for antioxidants from voltammetry
From the Table 2, it can be seen that the negative value of ΔG0 revealed the interaction process is spontaneous.
The following effects of variations in quinone structure on antioxidant activity can be deduced. Structure activity studies show that both the heterocyclic ring structure and the reactive orthoquinone moiety [49] of MD are responsible for higher antioxidant activity. According to previous reported data [50] if membered heterocyclic ring and orthoquinone moiety are present in a molecule, then antioxidant activity will be high. Since MD and MH molecules are having similar structural property, they have higher antioxidant capacity than others. As examination of Table 2 reflects MD shows the highest value of the binding constant (7.04 x 102) which ensures a stronger interaction with the radical at low concentration. Mansanone-D is having only CH3- group, so CH3- group is responsible for antioxidant activity of the compound. Similar behavior is observed for menadione compound [51]. MH is an sesquiterpenoid naphthoquinone with a structure similar to MD but the antioxidant activity of MH is low than MD, because different heterocyclic ring structure and substituted with a hydroxyl group at C-6 may not contribute to scavenge for DPPH radical. The antioxidant activity of phenolic hydroxyl in A ring is decreased significantly because of electrophylic effect of C ring [52]. In siss and 4MD molecules, radical scavenging activity is due to presence of electron donating substituent (-OCH3). However, it is lower than the MD and MH, because they are para quinones and also they do not have heterocyclic ring. The value of the change in free energy (ΔG0) is also following the similar trend MD>Siss>MH>4MD.
Conclusion
In this work, the electrochemical behavior of four quinones: MD, MH, Siss and 4MD and their interaction with DNA were investigated by CV technique. All the experimental results indicate that both electrostatic and intercalative binding modes were deduced from CV results, although intercalation mode of interaction of quinones-DNA complexes as the dominant mode. The strong interaction of MH with DNA was evident from their high binding constant values (i. e. 109 M-1). Interaction parameters like binding constant, binding site size, binding free energy and binding mode were determined. The negative values of the free energy indicated spontaneity of binding of the three quinones with DNA. These investigations showed that electrochemistry could provide a convenient way to characterize both the binding mode and the interaction mechanism of quinones with DNA, which is important for the design of new anticancer drugs. The antioxidant behavior and DNA binding studies were carried out for four quinones by CV technique and MD possess high antioxidant capacity(7.21 x 102)and MH has high DNA binding constant(1.04 x 109)(intercalation mode). All compounds shows antioxidant activity capacity and strong interaction with DNA.
Acknowledgement
The author Deepa would like to thanks the Basic Scientific Research (BSR), New Delhi for providing fellowship [No:UGC NO. F.4-/2006(BSR) 7-119/2007(BSR)] New Delhi, India.
References
- W. Zheng, S.Y. Wang, J. Agric. Food. Chem., 2001, 49, 5165-5170.
- Y.Z. Cai, M. Sun, H. Corke, J. Agric. Food Chem., 2003,51, 2288-2294.
- S. Ahmed, F. Shakeel, Czech J. Food. Sci., 2012, 30(2), 153-163.
- A.M. Pisoschi, G.P. Negulescu, 2011, 1(1), 1-10.
- S.T. Hussain, A. siddiqa, Europ. J. Chemistry., 2011,2(1), 109-112.
- Y.N. Ni, D.Q. Lin, S. Kokot, Anal. Chim. Acta.,2008, 606(1), 19-25.
- F. Araya, G. Huchet, I. McGroarty, G.G. Skellern, R.D. Waigh, Methods.,2007, 42(2), 141-149.
- J. Li, Y.L. Wei, L.M. Guo, C.H. Zhang, Y. Jiao, S.M. Shuang, C. Dong, Talanta., 2008, 76(1), 34-39.
- R.F. Pasternack, E.J. Gibbs, J.J. Villafranca, Biochem., 1983,22(10), 2406-2414.
- K.E. Erkkila, D.T. Odom, J.K. Barton, Chem. Rev., 1999, 99(9), 2777-2796.
- L.S. Ling, Z.K. He, F. Chen, Y.E. Zeng, Talanta.,2003, 59(2), 269-275.
- X.Q. He, Q.Y. Lin, R.D. Hu, X.H. Lu, Spectrochim. Acta A., 2007, 68(1), 184-190.
- E. Mirmomtaz, A.A. Ensafi, Electrochim. Acta., 2009, 54(18), 4353-4358.
- A.A. Ensafi, B. Rezaei, M. Amini, E.H. Bafrooei, Talanta., 2012,88, 244-251.
- C.L. Bian, Q.X. Zeng, L.J. Yang, H.Y. Xiong, X.H. Zhang, S.F. Wang, Sens. Actuators B.,2011, 156(2), 615-620.
- Y. Wang, Y. Ni, S. Kokot, Anal. Biochemistry., 2011, 419(2), 76-80.
- M.T. Cater, A.J. Bard, J. Am. Chem. Soc., 1987, 109(24), 7528-7530.
- X. Chu, G. Shen, J.H. Jiang, T.F. Kang, B. Xiong, R.Q. Yu, Anal. Chim. Acta., 1998, 373(1), 29-38.
- A.J. Lin, A.C. Sartorelli, J. Org. Chem.,1973, 38(4), 813-815.
- C. Asche, Mini- Reviews in Medicinal Chemistry.,2005, 5(5), 449-467.
- A.M.O. Brett, T.R.A. Macedo, D. Raimundo, M.H. Marques, S.H.P. Serrano, Biosens. Bioelectronics.,1998, 13(7-8), 861-867.
- D.T. Breslin, J.E. Coury, J.R. Anderson, L. Mc Fail-Isom, Y. Kan, L.D. Williams, L. Bottomley, G.B. Schuster, J. Am. Chem. Soc.,1997, 119 (21), 5043-5044.
- L.S. Lerman, J. Mol. Biol., 1961, 3(1), 18-30.
- E.L. Cavalieri, E.G. Rogan, Ann. N. Y. Acad., 2002, 959, 341-354.
- S. Rauf, J.J. Gooding, K. Akhtar, M.A. Ghauri, M. Rahman, M.A. Anwar, A.M. Khalid, J. Pharm. Biomed. Anal., 2005, 37(2), 205-217.
- Subhasini. K. Sripathy, Ph. D thesis, MKU, 1993.
- J. Johnson Inbaraj, R. Gandhidasan, R. Murugesan, Free Radic. Biol. Med., 1999, 26(9-10), 1072-1078.
- A. Shah, R. Qureshi, NK. Janjua, S. Haque, S. Ahmad, Anal. Sci., 2008, 24 (11), 1437-1441.
- P. SarathiGuin, S. Das, P.C.Mandal, Int. J. Electrochem., 2011, 2011, 1-22.
- J.Q. Chambers, S. Patai, Z. Rappoport, John Wiley & Sons Ltd., New York, 1974, 2, 737-791.
- M.O.F. Goulart, A.E.G. Sant’Ana, V. Horak, Micro-chim. Acta., 1986, 88(1-2), 23-26.
- M. Paulino, M. Hansz, N. Hikichi, G. Tabares, MP. Molina Portela, SH. Fernandez Villamil, CM. Sreider, AOM. Stoppani, An. Asoc. Quim. Argent, 1994, 82(5), 371-389.
- D.W. Pang, H.D. Abruna, Anal. Chem., 1998, 70 (15), 3162-3169.
- A. Shah, A. Rauf, A. Ullah, A. Munir, R. Qureshi, I. Ahmad, M.T. Soomro, Z. Rehman, J. Electrochem. Sci. Eng., 2013, 3(1), 19-27.
- M. Aslanoglu, Anal. Sci.,2006, 22(3), 439-443.
- X. Lu, M. Zhang, J. Kang, X. Wang, L. Zhuo, H. Liu, J. Inorg. Biochem., 2004, 98(4) 582-588.
- M.S. Ibrahim, Anal. Chimi. Acta., 2001, 443(1), 63-72.
- D. Xu, X. Wang, L. Ding, Carbohydr. Polym., 2011, 83(3), 1257-1262.
- M.T. Carter, M. Rodriguez, A.J. Bard, J. Am. Chem. Soc., 1989, 111(24), 8901-8911.
- M. Aslanoglu, C.J. Isaac, A. Houlton, B.R. Horrocks, Analyst., 2000, 125(10), 1791-1798.
- A. Shah, R. Qureshi, A.M. Khan, R.A. Khera, F.L. Ansari, J. Braz. Chem. Soc., 2010, 21 (3) 447-451.
- Z. Xu, G. Bai, C. Dong, Bioorg. Med. Chem.,2005, 13(20), 5694-5699.
- B.L. Funt, D.G. Gray, Can. J. Chem.,1968, 46, 1337-1340.
- M.K. Kalinowski, J. Klimkiewicz, Monatsh. Chem.,1983, 114(10), 1035-1043.
- A.M. Pisoschi, M.C. Cheregi, A.F. Danet, Molecules,2009, 14(1), 480-493.
- E.I. Korotkova, Y.A. Karbainov, A.V. Shevchuk, J. Electroanal. Chem., 2002, 518(1), 56-60.
- S. Chevion, M.A. Roberts, M. Chevion, Free Radical. Biol. Med., 2000, 28(6), 860-870.
- B. Yang, A. Kotani, K. Arai, F. Kusu, Anal. Sci.,2001, 17(5), 599-604.
- M.P. Nambiar, R. Murugesan, H.C. Wu, J. Cell. Physio.,1998, 176(1), 40-49.
- K. Mukai, K. Daifuku, K. Okabe, T. Tanigaki, K. Inoue, J. Org. Chem.,1991, 56(13), 4188-4192.
- S. Kumar, S. Gautam, A. Sharma, Mutation Research, 2013, 755(1), 30-41.
- C.Z. Lin, C.C. Zhu, M. Hu, A.Z. Wu, B.Z. DawA, K.S. Qimei, J. Intercult. Ethnopharmacol.,2014, 3(3), 123-127.