Research Article - Der Pharma Chemica ( 2018) Volume 10, Issue 2
Interaction of Azo Dyes with BSA and Adenine: Spectral, Electrochemical and Molecular Docking Methods
Rajendiran N*, Thulasidhasan J and Suresh M
Department of Chemistry, Annamalai University, Annamalai Nagar-608 002, Tamil Nadu, India
- *Corresponding Author:
- Rajendiran N
Department of Chemistry
Annamalai University
Annamalai Nagar-608 002, Tamil Nadu, India
Abstract
Interaction of Azobenzene (AB), 4-Hydroxyazobenzene (HAB) and 2,4-dihydroxyazobenzene (or) Sudan Orange G (SOG) with BSA and DNA base (adenine) were investigated by UV-Visible, fluorescence and ATR-IR spectroscopy, cyclic voltammetry and molecular docking methods. With on addition of azo dyes into the BSA and adenine molecules different emission spectral shifts observed, i.e. (i) Upon increasing the concentration of AB molecule, red shifted quenching is noticed in the BSA molecule (338-367 nm), (ii) No significant spectral shifts noticed in the HAB molecule and (iii) Blue shift (338-320 nm) is observed in the SOG molecule. Azo dyes can bind to BSA with stoichiometric ratio of 1:1 and the protein-ligand complexes are stabilized mainly by hydrophobic and van der Waals interaction. IR spectral analysis shows, the azo group (N=N) stretching frequency for AB, HAB and SOG molecules were shifted in the BSA complex. In cyclic voltammetry, the concentration of the dyes added to the BSA solution, a reductive peak appeared in AB, but this peak is not noticed in HAB and SOG. Molecular docking studies specify that the dyes are located within the binding pocket of sub-domain IIIA in BSA and hydrophobic force play a major role in the binding. As a consequence of this study, the addition of a hydroxyl group to the aromatic ring resulted in a decrease in the binding affinity.
Keywords
Azo dyes, Bovine serum albumin, Fluorescence quenching, Cyclic voltammetry, Molecular docking
Introduction
Dyes are being increasingly used for clinical and medicinal purposes [1]. The discovery that some dyes would stain certain tissues and not others led to the idea that dyes might destroy pathogenic organisms without causing appreciable harm to the host. Azo dyes are fat soluble, powdered with orange or red colour appearances. The additive is mainly used to colour waxes, oils, petrol, solvents and polishes. Laboratory tests of azo dyes on rats showed growth of cancerous tumors in the liver and bladder [2,3]. Although the use of azo dyes in foods is now banned in many countries because they have been classified as category 3 carcinogens by the International Agency for Research on Cancer, these dyes have been adopted for colouring a variety of foods (Foodstuffs); including particular brands of curry and chilli powder [4-6]. In recent years, events impacting on the security of food supply have caused wide food scare around the world.
Serum albumins are the major soluble protein and it has many physiological functions. They play a dominant role in transport and disposition of various dyes and drugs in blood. The absorption, distribution, metabolism, excretion properties as well as the stability and toxicity of dyes and drugs can be significantly affected as a result of their binding to serum albumins [7]. Further, there is evidence of conformational changes of serum albumin induced by its interaction with low molecular weight dyes and drugs, which appears to affect the secondary and tertiary structure of albumins [8]. Consequently, the investigation on the interaction of dyes and drugs to serum albumin is of great importance.
BSA was selected as our protein model because of its low cost, ready availability, unusual ligand-binding properties, and the results of the studies are consistent with the fact that bovine and human serum albumins are homologous proteins [9-11]. Bovine Serum Albumin (BSA) is made up of three homologous domains (I, II, III); each domain in turn is the product of two sub-domains. And BSA has two tryptophans, Trp- 212, which is located within a hydrophobic binding pocket of the protein, and Trp-134, locating on the surface of the molecule [12], domains (I, II, III) that are divided into nine loops (L1-L9) by 17 disulfide bonds. The loops in each domain are made up of a sequence of large-small-large loops forming a triplet. Each domain in turn is the product of two sub-domains (IA, IB, etc.,). This kind of study may provide salient information on the structural features that determine the therapeutic effectiveness of drugs/dyes, and hence become an important research field in chemistry, life sciences and clinical medicine [13,14].
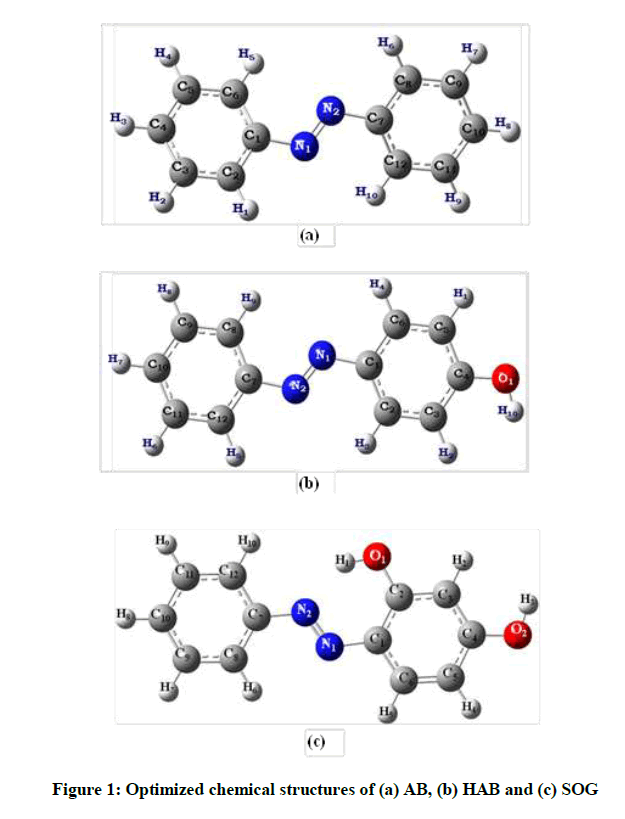
Thus, providing insights on the interaction between azo dyes and bio-macromolecules, including proteins, is a necessity. This approach originally developed to illuminate the structure and excited state behavior of organic molecules and dyes included in cyclodextrin macrocycles through experimental and theoretical [15-20] explorations. In view of increasing attention directed toward the importance of investigating interaction between proteins and drugs/dyes, we present in this work utilizing UV-Visible and fluorescence spectroscopy, cyclic voltammetry and molecular docking studies for investigating the interaction between three azo dyes, namely-Azobenzene (AB), 4-Hydroxylazobenzene (HAB) and 2,4-Dihydroxyazobenzene (or) Sudan Orange G (SOG) (Figure 1) with BSA/adenine. In particular, our investigation is focused on fluorescence quenching, determination of binding constant and binding sites of the dyes with BSA/adenine system, energy transfer and binding distance between BSA/adenine and drugs, thermodynamic free energy and binding modes and conformation changes of BSA/adenine upon binding to drugs.
Experimental Section
Materials
AB, HAB, SOG, adenine and BSA were purchased from Sigma-Aldrich chemical company, USA, and used without further purification. The purity of the compound was checked for similar fluorescence spectra when excited with different wavelengths.
Preparation of experimental solution
The concentration of stock solution of the BSA and adenine was 2 × 10-3 M. The stock solution (0.2 ml) was transferred into 10 ml volumetric flasks. To this, varying concentration of above dye solution (1.0 × 10-3 to 1.0 × 10-2 M) was added. The aliquot solution was diluted to 10 ml with triply distilled water and shaken thoroughly. The final concentration of BSA and adenine in all the flasks was 2 × 10-5 M. All solutions were stored in a refrigerator at 4°C in the dark.
Instruments
Spectral measurements (Absorbance and fluorescence)
UV-visible absorption spectra were recorded using a Shimadzu UV 2600PC model equipped with 1.0 cm quartz cells at 25°C. Buffer (control) and samples were placed in the reference and sample cuvetes respectively. Fluorescence measurements were performed using a Shimadzu spectrofluorimetry model RF-5301 equipped with a 1.0 cm quartz cell. The excitation and emission slit widths were fixed at 5 or 10 nm to the corresponding azo dyes. The excitation wavelength was set at 280 nm (Excitation of the Trp and Tyr), and the emission spectra were read at 290- 500 nm at a scan rate of 100 nm/min-1. Experiments were carried out at room temperatures (298 K) with recycle water keeping the temperature constant.
Electrochemical measurements (Cyclic voltammetry)
Cyclic voltammetry measurements were performed through an electrochemical workstation (model-CHI 620D, CH Instruments, USA) with a three electrode system: surface area 0.1963 cm2 platinum disc as working electrode, saturated silver electrode as reference electrode and a platinum foil as counter electrode. Prior to use, the working electrode was polished with 0.05 μm alumina and thoroughly washed in an ultrasonic bath for 5 min. Before experiments, the solution within a single-compartment cell was deaerated by purging with pure N2 gas for 5 min.
Molecular docking studies
Molecular docking calculations were carried out using online docking server (http://www.dockingserver.com) [21]. The MMFF94 force field was used for energy minimization of the azo dyes (AB, HAB and SOG) molecules using Docking Server [22]. Gasteiger partial charges were added to the drug atoms. Non-polar hydrogen atoms were merged, and rotatable bonds were defined. Docking calculations were carried out on 4F5S protein model. Essential hydrogen atoms, Kollman united atom type charges, and solvation parameters were added with the aid of Auto Dock tools [23]. Affinity (grid) maps of 20 × 20 × 20 Å grid points and 0.375 Å spacing were generated using the Auto grid program [23]. Auto Dock parameter set- and distance-dependent dielectric functions were used in the calculation of the van der Waals and the electrostatic terms, respectively. Docking simulations were performed using the Lamarckian genetic algorithm (LGA) and the Solis & Wets local search method. Initial position, orientation, and torsions of the drug molecules were set randomly. The population size was set to 150. During the search, a translational step of 0.2 Å, and quaternion and torsion steps of 5 were applied.
Results and Discussion
Absorption spectral measurement
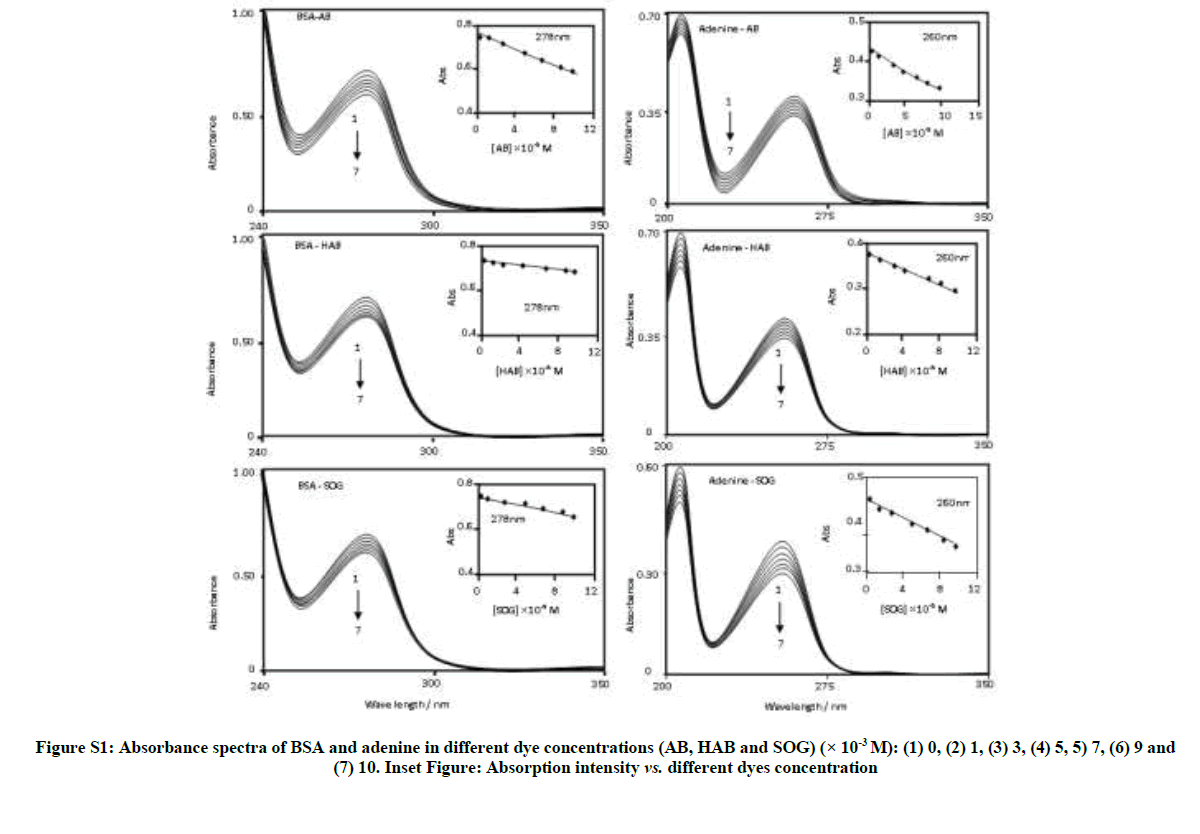
Table S1 and Figure S1 depict the absorption spectra of BSA and adenine in aqueous solution containing different concentrations of AB, HAB and SOG. In the non-existence of BSA and adenine, the above azo dyes shows the absorption maxima at ~319, 228 nm for AB, ~342, 228 nm for HAB and ~382, 249 220s for SOG, respectively [15]. In aqueous solution the absorption maximum of BSA observed at ~280 nm and adenine appears at ~260 nm (Figure 2). With an increasing the concentration of azo dyes, the absorbance maxima of BSA and adenine decreased regularly (Figure S1). No notable spectral shift observed in BSA and adenine with varying the dye concentrations except a decrease in the absorbance of BSA and adenine. The absorbance of the solutions recorded after 12 hours remains unchanging indicating that these azo dye molecules are interacted with BSA and adenine solution without decaying on storage due to the formation of protein-ligand (dye) complex. The measured absorbance, plotted against the dye concentration is shown in the insert of Figure S1. The absorbance decreases in the azo dyes indicate the interaction of these molecules in the amino acid residues in BSA and adenine molecules [24-26].
| Azo dyes (M) × 10-5 |
BSA | Adenine | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AB | HAB | SOG | AB | HAB | SOG | |||||||||||||
| λabs | logε | λflu | λabs | logε | λflu | λabs | logε | λflu | λabs | logε | λflu | λabs | logε | λflu | λabs | logε | λflu | |
| 0 (Without dye) |
278 | 4.56 | 338 | 278 | 4.54 | 338 | 278 | 4.54 | 338 | 260 | 4.31 | 302 326 420 |
260 | 4.29 | 302 | 260 | 4.29 | 302 324 419 |
| 1 | 278 | 4.55 | 344 | 278 | 4.47 | 338 | 278 | 4.47 | 335 | 260 | 4.31 | 302 326 |
260 | 4.29 | 300 334 |
260 | 4.29 | 302 321 |
| 3 | 278 | 4.54 | 351 | 278 | 4.43 | 338 | 278 | 4.43 | 334 | 260 | 4.28 | 302 346 |
260 | 4.28 | 300 334 |
260 | 4.28 | 303 321 |
| 5 | 278 | 4.53 | 360 | 278 | 4.4 | 338 | 278 | 4.4 | 329 | 260 | 4.26 | 302 361 |
260 | 4.28 | 300 333 |
260 | 4.28 | 303 321 |
| 7 | 278 | 4.52 | 362 | 278 | 4.39 | 338 | 278 | 4.39 | 327 | 260 | 4.25 | 301 363 |
260 | 4.27 | 301 332 |
260 | 4.27 | 304 321 |
| 9 | 278 | 4.52 | 365 | 278 | 4.37 | 337 | 278 | 4.37 | 323 | 260 | 4.23 | 301 364 |
260 | 4.26 | 301 334 |
260 | 4.26 | 304 320 |
| 10 | 278 | 4.51 | 367 | 278 | 4.35 | 337 | 278 | 4.35 | 320 | 260 | 4.22 | 301 364 |
260 | 4.24 | 302 335 |
260 | 4.24 | 303 320 |
| Excitation wavelength (nm) | - | 280 | - | 280 | - | 280 | 270 | 270 | 270 | |||||||||
| R0 (nm) | - | 0.68 | - | 0.88 | - | 0.98 | 0.36 | 0.47 | 0.37 | |||||||||
| J × (10-14) [cm3 L mol-1] |
- | 1.68 | - | 1.82 | - | 1.96 | 0.85 | 0.87 | 0.91 | |||||||||
| r (nm) | - | 0.73 | - | 0.69 | - | 0.53 | 1.18 | 1.11 | 1.23 | |||||||||
| E | 0.28 | 0.22 | 0.23 | 0.25 | 0.24 | 0.21 | ||||||||||||
AB-Azobenzene, HAB-Hydroxy azobenzene, SOG-Sudan orange G
Table S1: Absorption and fluorescence maxima of BSA and adenine with different AB, HAB and SOG concentrations
Fluorescence spectral measurement
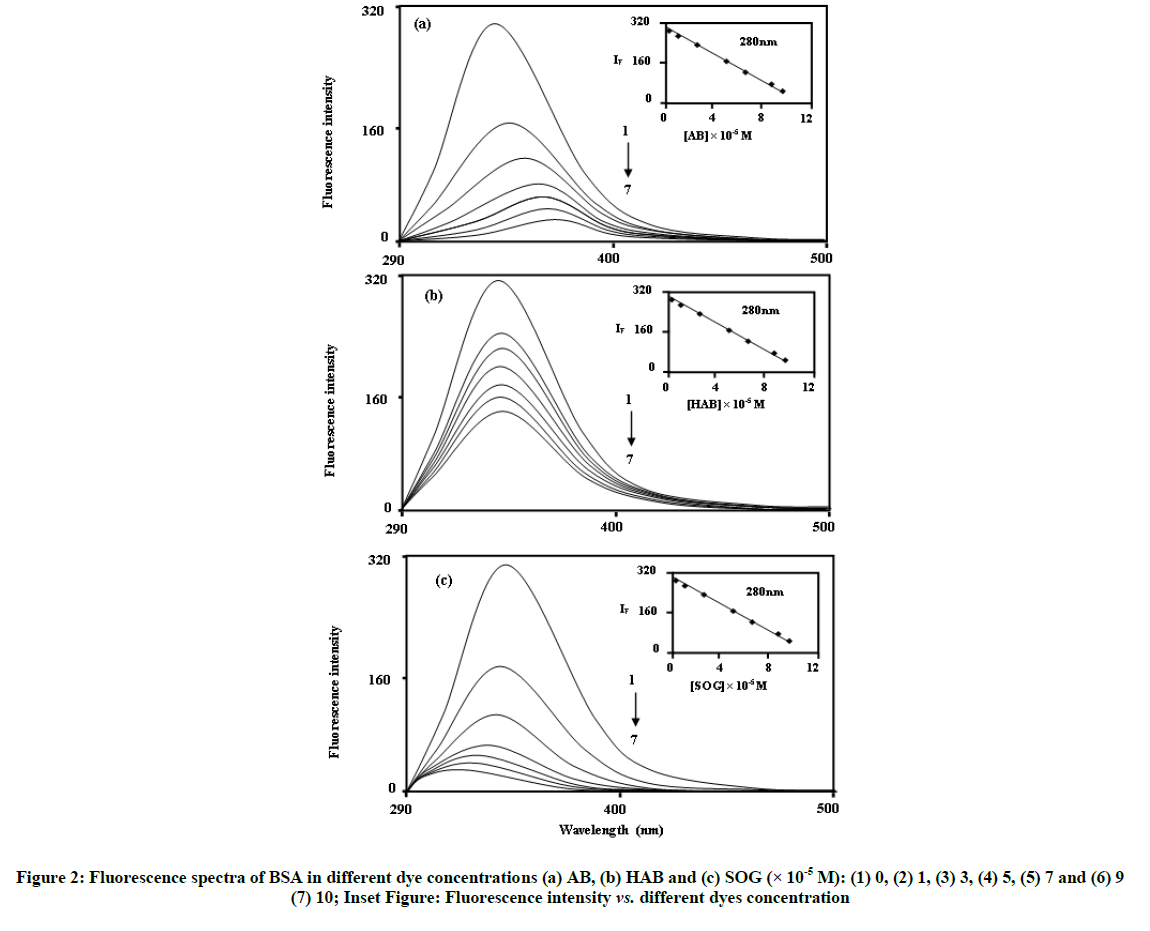
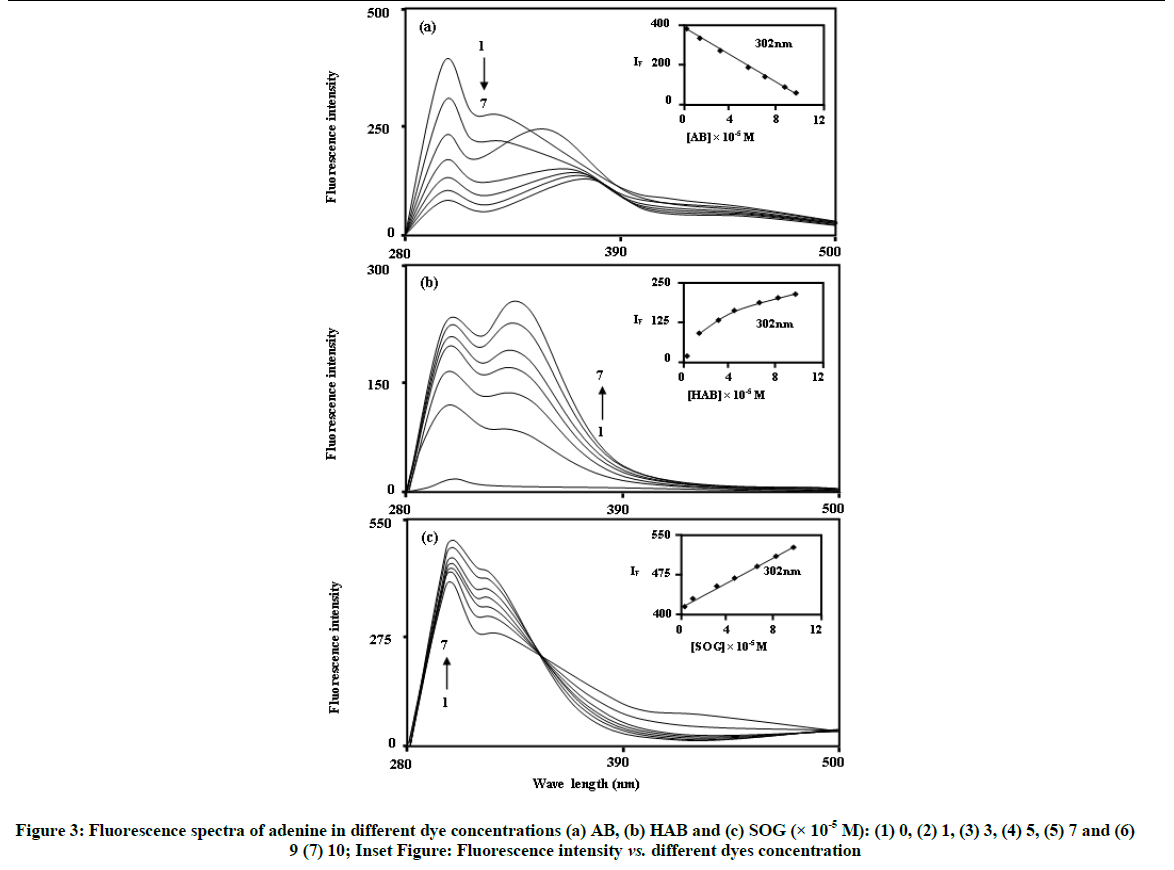
Figure 3 shows the fluorescence spectra of BSA and adenine in different concentrations of AB, HAB and SOG. BSA shows a strong fluorescence emission band at ~340 nm when the excitation wavelength is fixed at 280 nm. In the absence of BSA and adenine, azo dyes show the emission maxima at ~370 nm for AB, ~390 nm for HAB and ~430 nm for SOG respectively [15]. In BSA and adenine solutions, the fluorescence spectra of the above azo dye molecules are more sensitive than the absorption spectra. Interestingly, with on addition of azo dyes into the BSA molecule different spectral shifts observed; i.e. (i) Upon increasing the concentration of AB molecule, red shifted quenching is noticed in the BSA molecule (338 to 367 nm), (ii) No significant spectral shifts noticed in the HAB molecule and (iii) Blue shift (338 to 320 nm) is observed in the SOG molecule (Figure 3). The above results indicate that BSA binds to the azo dyes in different manner. The large red shift observed with increasing AB concentration, suggest that the fluorophore of BSA is placed in a more hydrophilic environment after the addition of AB [27]. In SOG, the blue shifted emission wavelength suggests a decrease in the polarity of the microenvironment around fluorophore [28].
In the absence of azo dyes, the fluorescence spectra of adenine excited at 270 nm with slit width 10 nm, consists of three distinct bands at around 302 nm (F1) 326 nm (F2) 420 nm (F3). The intensity of the bands is in the order F1 > F2 > F3. The yields of the bands are in conformity with some other findings for similar other compounds. Figure 3 shows that the fluorescence intensity of adenine decreased as the concentration of AB increased. However, with the increasing concentration of HAB and SOG, the fluorescence intensity of adenine increased.
The interaction of the above dyes with Cyclodextrin (CD) is given below. The absorption and fluorescence characteristics of the azo dyes are seen to undergo drastic changes in the presence of cyclodextrin [15-20]. As mentioned earlier, in the presence of BSA/adenine, the absorption and emission maxima of these dyes are decreased. However, upon increasing the CD concentration, the fluorescence intensity of the AB molecule increases considerably at the same emission wavelength (370 nm) whereas the emission maxima of HAB and SOG azo dyes are largely red shifted from 390-410 nm to ∼458 nm and the emission intensity increases with increasing the CD concentrations. The substantial increase in the fluorescence intensity compared with absorbance shows that the quantum yields of both molecules increase in the presence of the CD. The above results show that the interaction of the azo dyes with the CD is different from BSA/adenine. This is because non-polar interactions are predominant in the CD cavity while polar interactions play major role in the BSA/adenine.
Quenching mechanism
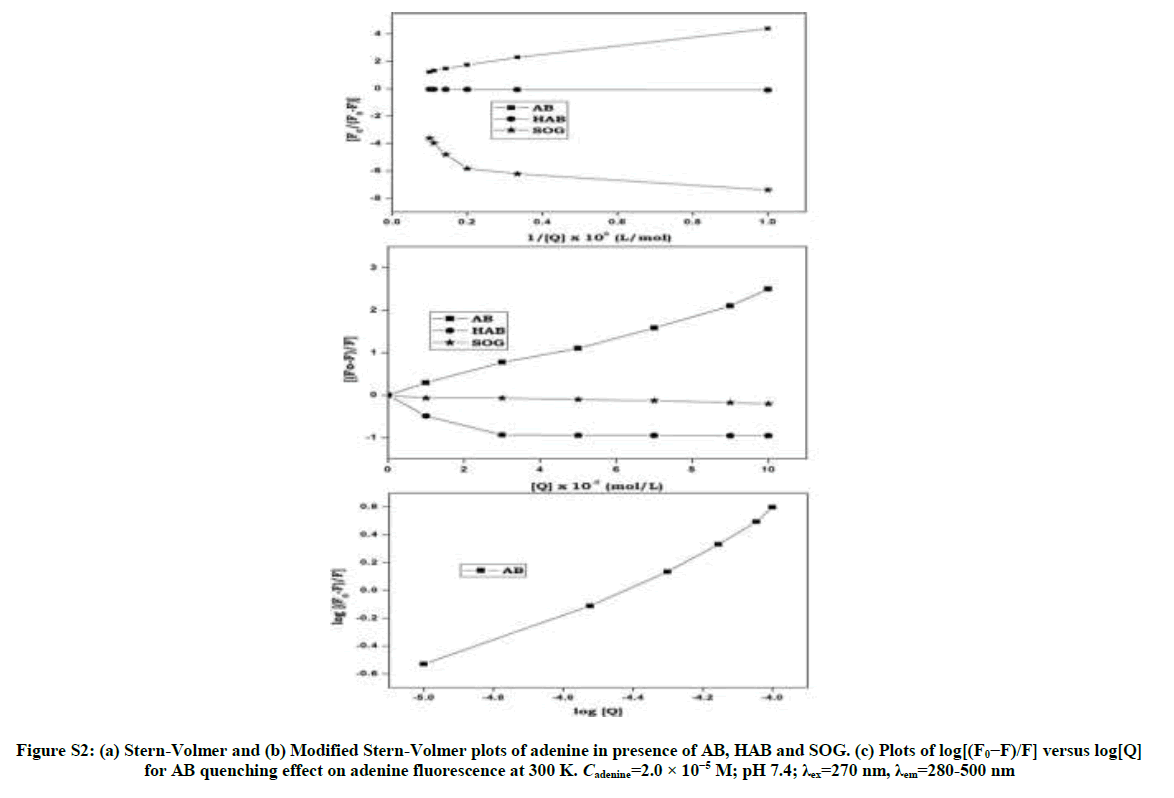
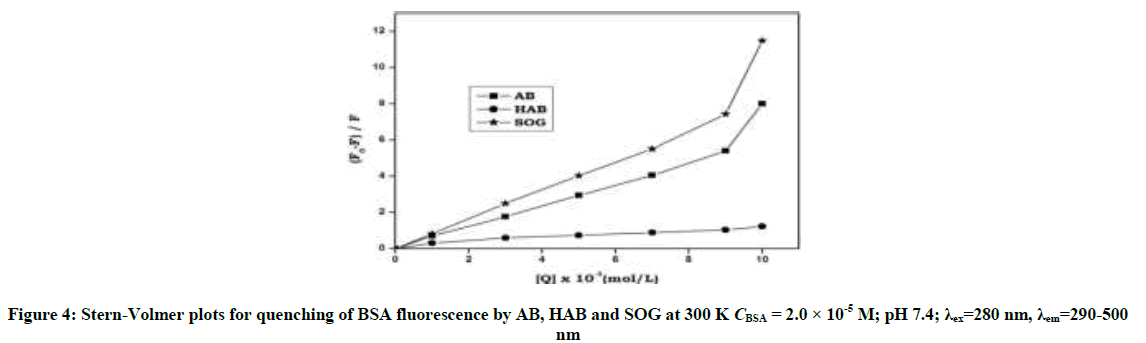
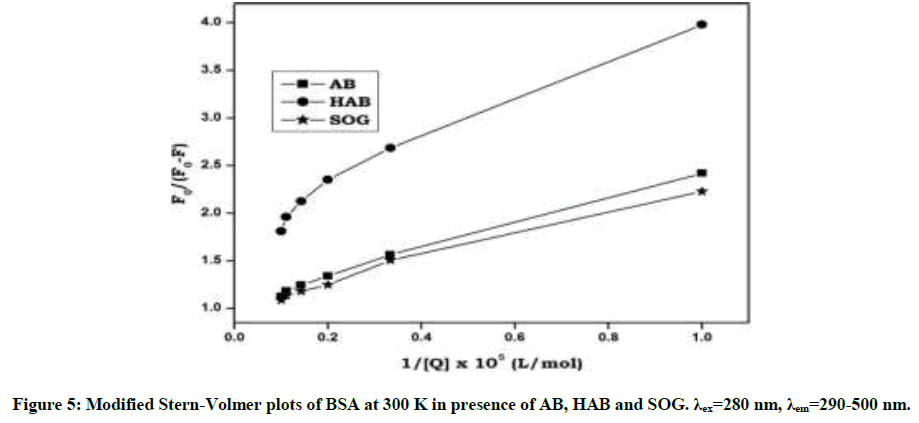
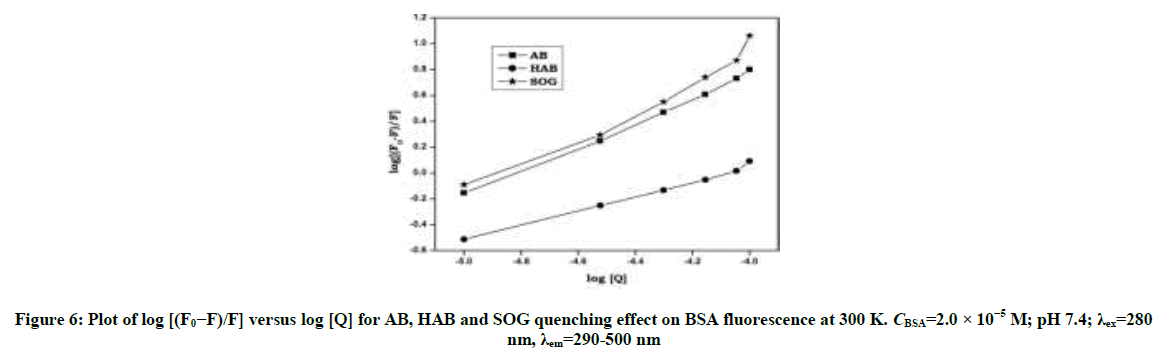
Upon addition of azo dyes, the fluorescence intensity of BSA and adenine was quenched. The fluorescence quenching data were analyzed by Stern-Volmer [29,30] Equation: (S1 and S2) Stern-Volmer plots for the quenching of BSA and adenine by the azo dyes at 300 K are shown in Figure 4 and Figure S2. The curves are linear with positive deviation and the calculated quenching constants of the corresponding dyes are listed in Table 1. It is known that linear Stern-Volmer plots represent a single quenching mechanism, either static or dynamic [31]. The maximum scatter collision quenching constant of various quenchers with biopolymer is 2.0 × 1010 M-1 s-1 [32]. In the present study, the values of the rate constant kq (KSV/τ0) for the quenching of BSA caused by the azo dye compounds are all much larger than the maximum diffusion collision quenching rate constant for the scatter mechanism. Hence, we propose that the quenching is not initiated by the dynamic collision, but originates from the formation of a complex. If the resulting plots exhibit a good linear relationship, this is generally indicative of the purely collisional quenching process or static quenching process. However, the Stern-Volmer plot showed a positive deviation (Figure 4) indicates the presence of both static and dynamic quenching [33]. Therefore, the data were processed based on a following modified Stern-Volmer Equation [34].
| Dyes | Ksv (105 M-1) | Kq (1013 M-1s-1) | Ra | SD | ||
|---|---|---|---|---|---|---|
| S-V quenching | BSA | AB | 5.51 | 5.51 | 0.998 | 0.492 |
| HAB | 1.45 | 1.45 | 0.996 | 0.668 | ||
| SOG | 7.19 | 7.19 | 0.997 | 0.592 | ||
| Adenine | AB | 2.75 | 2.75 | 0.999 | 0.761 | |
| HAB | -2.49 | -2.49 | 0.998 | 0.592 | ||
| SOG | -2.49 | -2.49 | 0.998 | 0.761 | ||
| Modified S-V quenching |
BSA | Drugs | Ka (104 M-1) | ΔG0 (kJ/mol) | n | Ra |
| AB | 2.11 | -21.56 | 0.992 | 0.995 | ||
| HAB | 1.45 | -20.87 | 0.962 | 0.998 | ||
| SOG | 2.18 | -20.61 | 1.024 | 0.997 | ||
| Adenine | AB | 1.44 | -20.98 | 0.935 | 0.996 | |
| HAB | -5.96 | -17.46 | - | 0.998 | ||
| SOG | -3.71 | -17.46 | - | 0.999 |
Ra-Linear correlation coefficient, SD-Standard deviation, n-Number of binding site
Table 1: Stern-Volmer quenching constant (Ksv) and Modified Stern-Volmer association constant Ka and bimolecular quenching rate constant (Kq) of the BSA and adenine - AB, HAB and SOG systems at temperature 300 K; λex=280 nm, λem=290-500 nm, (pH-7.4)
Stern-Volmer Equation
F0/F = 1 + KSV [Q] (S1)
Where, F0 and F are the steady-state fluorescence intensities in the absence and presence of quencher (drug), KSV is the Stern-Volmer quenching constant and [Q] is the concentration of quencher (SP or SD). The quenching rate constants (Kq) were evaluated using the Equation:
KSV = kqτ0 (S2)
Where, τ0 is the average lifetime of the protein without the quencher.
log [(F0–F)/ F]=log Ka + n log [Q] (S3)
Where, Ka is the binding constant and n is the number of binding sites, F0 is the fluorescence intensity of free BSA/adenine and F is the consecutive fluorescence on addition of the sulpha drugs. Plot of log [(F0– F)/F] against log [Q] was used to determine the values of K and n from the intercept and slope respectively. R0 can be calculated from donor emission and acceptor absorption spectra using the Forster formula:
E =1- F/F0 = R0/(R06+r6) (S4)
R06 = 8.79 × 10-25 K2 n-4 φ J (S5)
 (S6)
(S6)
From Equation (S4)-(S6), r is the distance between the acceptor and the donor, and R0 is the Forster critical distance, at which 50% of the excitation energy is transferred to the acceptor. K2 is the spatial orientation factor of the dipole, n is the refractive index of the medium, J is the overlap integral of the fluorescence emission spectrum of the donor and the absorption spectrum of the acceptor, F(λ) is the fluorescence intensity of the fluorescent donor at wavelength k and ε (λ) is the molar absorptivity of the acceptor at wavelength (λ).
The binding constant Ka and binding sites n can be obtained by using fluorescence intensity data. When azo dyes bind independently to a set of equivalent sites on a protein molecule the equilibrium between free and bound molecules (interaction complex) is calculated from the equation (S3). The values of Ka and n were obtained from the intercept and slope of the plot of log[(F0-F)/F] vs. log[Q] (Figures S3 and S4). The values of Ka were found to be (2.11 × 105, 1.45 × 105 and 2.18 × 105 l/mol-1) and those of n were noticed to be 0.99, 0.96 and 1.02 respectively (Table 1). The values of binding sites close to unity indicated that there was only one independent class of binding site on BSA for dyes. AB molecule has high Ksv and Ka values with BSA and adenine than other dyes indicates AB molecule has more quenching behavior with BSA/adenine molecule than HAB and SOG (Table 1).
Thermodynamic parameters and nature of the binding forces
Intermolecular hydrogen bonding, electrostatic forces, hydrophobic forces, and van der Waals interactions are essentially four types of noncovalent interactions that could play a major role in small organic molecules (Dyes) binding to biological macromolecules (Proteins) [35]. Sign and magnitude of the thermodynamic parameters are important information for confirming the main forces involved in the binding reaction. For this purpose, thermodynamic parameters (ΔG0) were calculated from the Equation 1 and it is summarized in Table 1.
ΔG0 = - RT ln Ka (1)
Where, R is the universal gas constant (8.314 J/mol-1/K-1), T is Temperature (K) and Ka is the binding constant obtained from Equation (S2). The negative ΔG0 values showed the spontaneous nature of the binding process.
Energy transfer from BSA to azo dyes
Fluorescence Resonance Energy Transfer (FRET) is a dependable method for studying protein–ligand interactions and estimation of the distance between the dyes and tryptophan residues of the protein. The energy transfer efficiency (E) is defined by the Equation (S4).
On the basis of equations (S4)-(S6), the parameters (J, R0, E and r) of azo dyes with BSA and adenine are listed in Table S1. In general, the average distances between a donor and acceptor fluorophore after interaction between these dyes with BSA/adenine is less than 8 nm, and 0.5 R0 < r < 1.5 R0 [36] these accord with the conditions of Fosters non-radiative energy transfer theory indicating the static quenching interaction between BSA/adenine with AB, HAB and SOG. The results in Table S1 indicate that, the energy transfer efficiency (E) of AB with BSA and adenine has more than that of HAB and SOG with BSA/adenine.
ATR-IR spectral studies
IR spectra of BSA, azo dyes and BSA with azo dyes are shown in Figure 5. Changes in spectral shape and band position of pure BSA, AB, HAB and SOG in the corresponding interactions confirm the formation of the protein-dye complexes. Azo group (N=N) stretching frequency for AB, HAB and SOG appeared at 1453, 1450 and 1458 cm-1 was moved in the BSA complex to 1453, 1466 and 1449 cm-1 respectively. The C-N stretching frequency appeared at 1658, 1643 and 1626 cm-1 for AB, HAB and SOG was moved in the BSA complex to 1647, 1643 and 1649 cm-1 respectively. In HAB and SOG, the –OH stretching vibration frequency is appeared at 3355 and 3266 cm-1 is moved to lower and higher frequencies in the BSA: azo complex (3285 and 3284 cm-1).
Amide I band shifted from 1649 cm-1 to 1647, 1643 and 1649 cm-1 in AB, HAB and SOG molecules with BSA (Figure 5). The amide II band shifted from 1545 to 1583 cm-1 in AB-BSA and it does not appear in the BSA with HAB and SOG molecules. The above results indicated the change in secondary structure of BSA upon binding to the dyes. The different IR spectrum obtained for pure BSA, dyes and BSA-dye complex conform the interactions present between the BSA and azo dyes.
Cyclic voltammetry
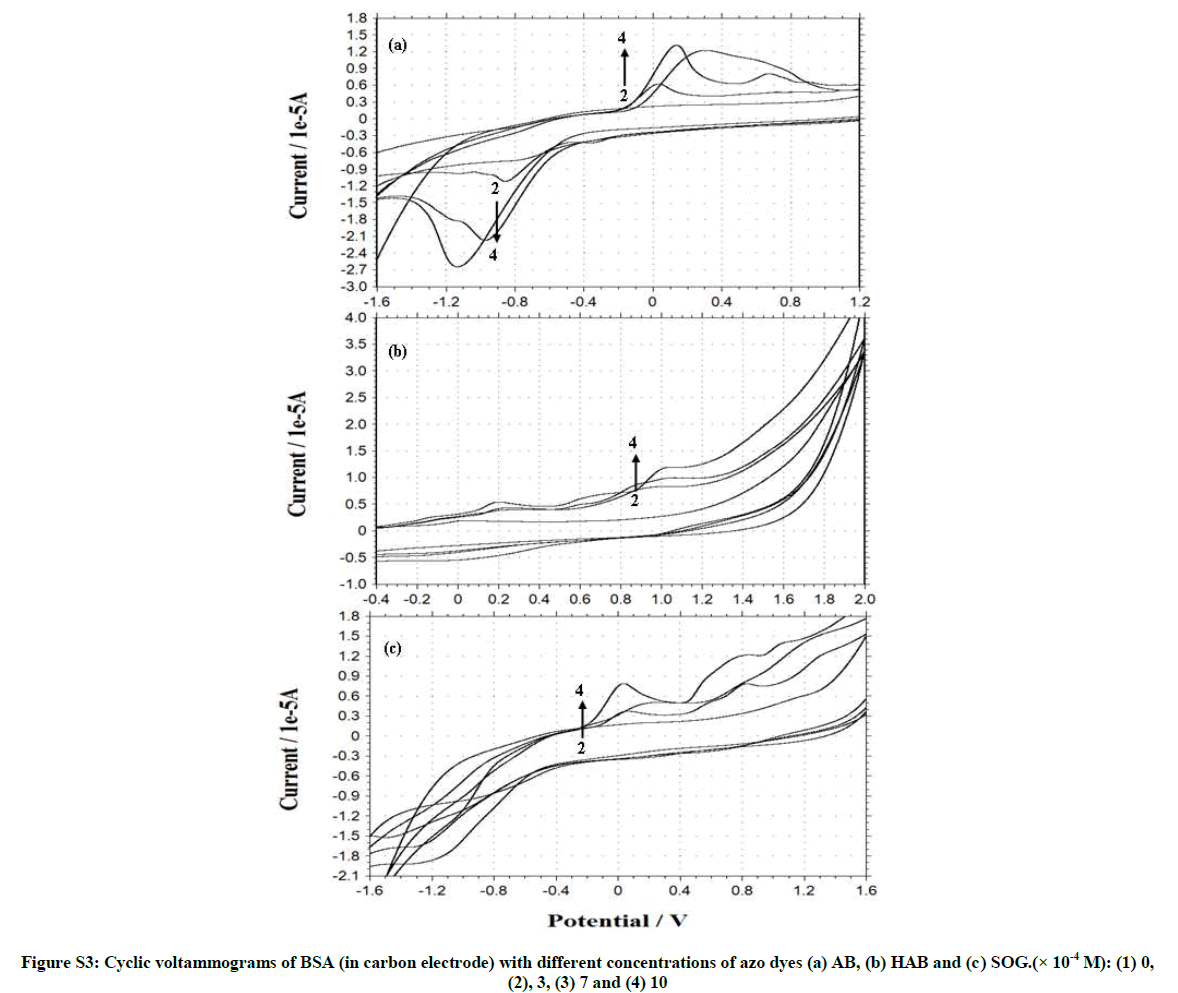
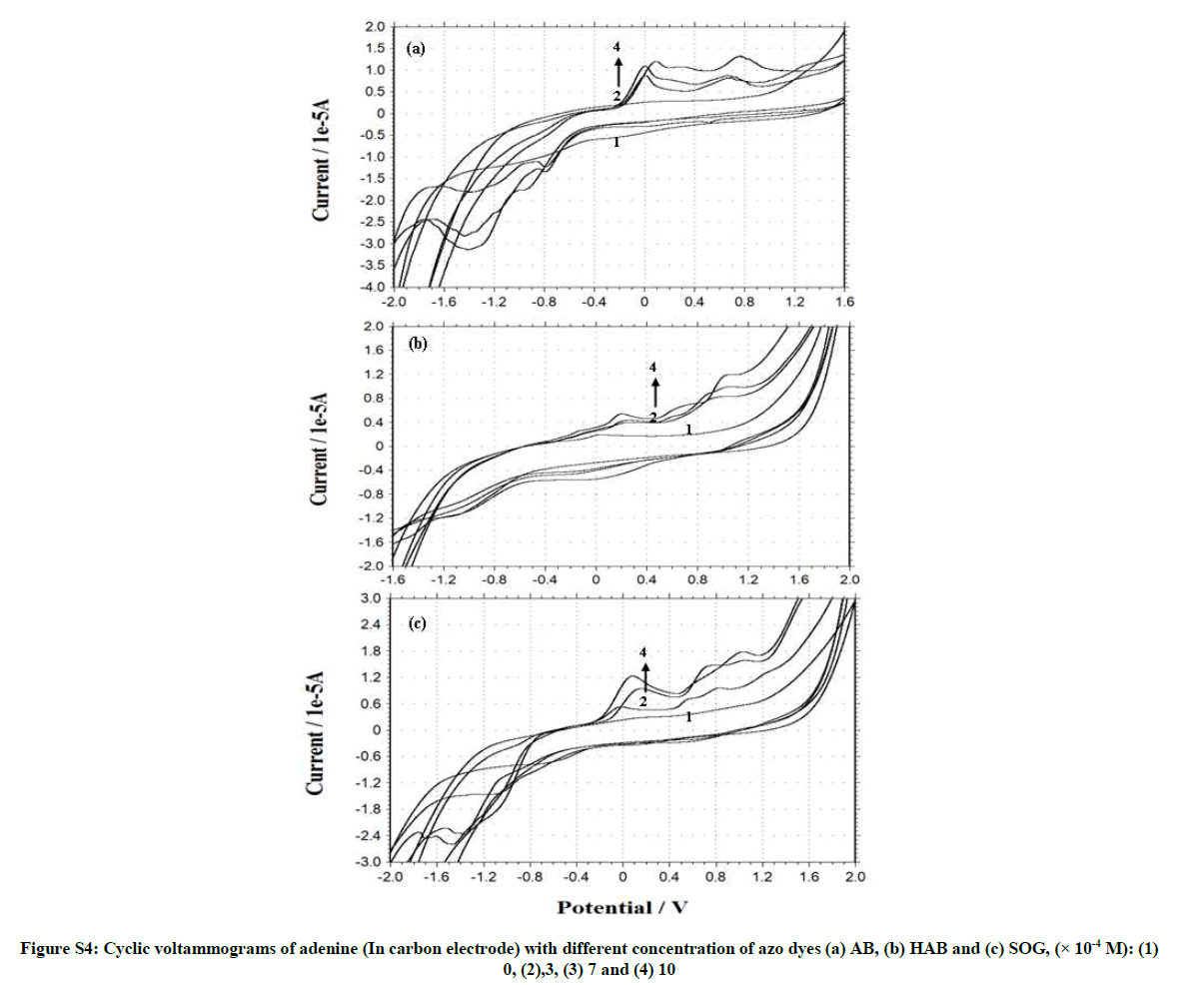
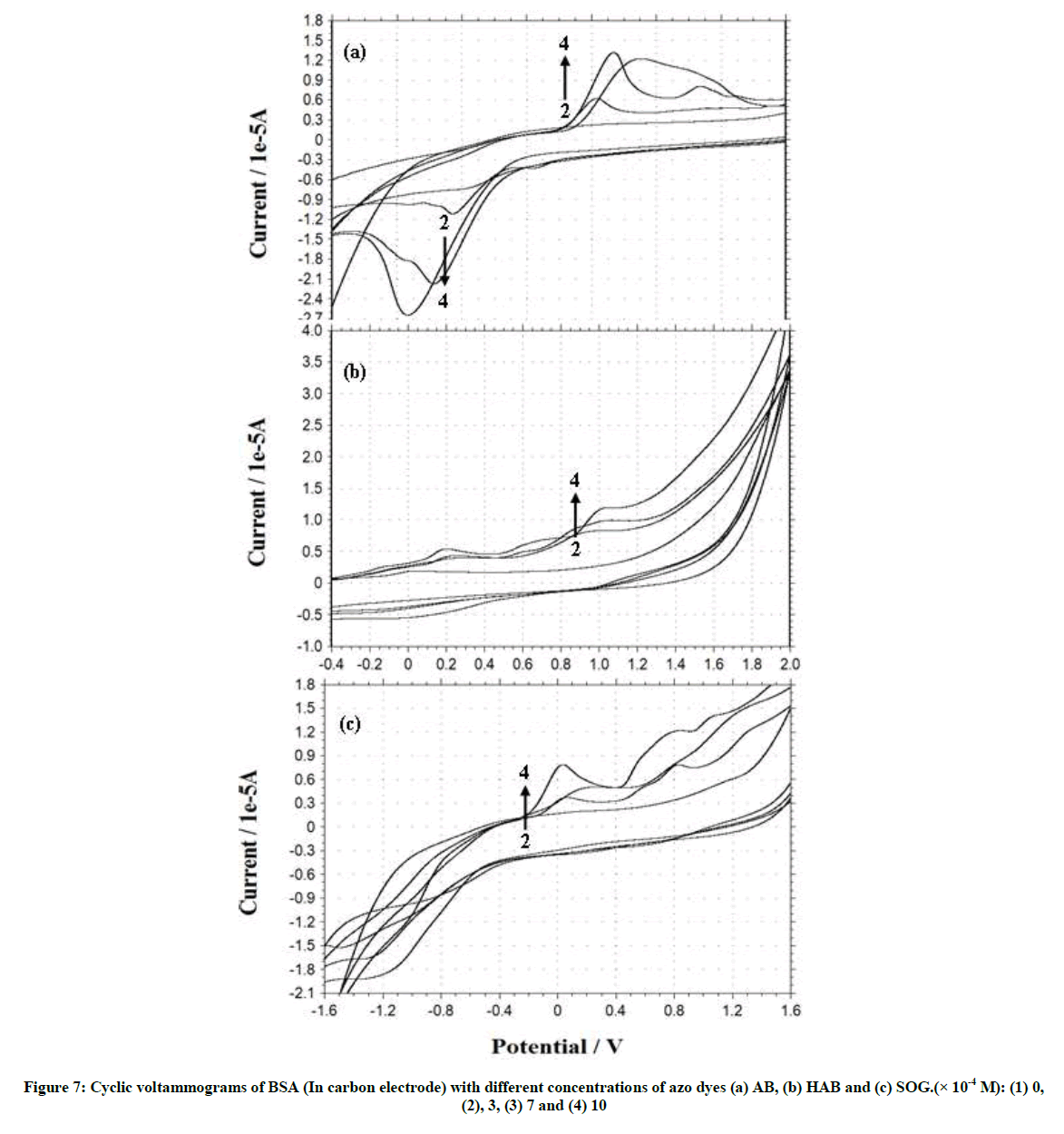
Figures S3 and S4, Tables 2 and 3 shows the cyclic voltammogram of BSA and adenine in the absence and presence of different amounts of AB, HAB and SOG. During the experimental process, BSA and adenine concentration were kept constant (2.0 × 10-6), and the concentrations of the dyes were varied from 1.0 × 10-3 M to 1.0 × 10-2 M. AB, HAB and SOG have a oxidative peak at 0.573 V, 0.714 V and 0.354 V respectively (Figures S3 and S4), indicated that the electrochemical behavior of the dyes on glassy carbon electrode was irreversible in pH ~7.4 buffer solution. When the concentration of the dyes added to the BSA solution, a reductive peak appeared in AB (Figure S3a), but this peak is not noticed in HAB and SOG (Figures S3b and c).
| Dye-BSA | Dye concentration (× 10-4) |
Epa | Ipa | Epc | Ipc | Epa-Epc/2 | Ipa/Ipc |
|---|---|---|---|---|---|---|---|
| AB | 8 × 10-5 | -1171 | -1.345 | 117 | 0.573 | -644 | -2.347 |
| HAB | 8 × 10-5 | - | - | 889 | 0.714 | -444 | - |
| SOG | 8 × 10-5 | - | - | -40 | 0.354 | 20 | - |
| BSA only | 2 × 10-5 | - | - | 973 | 0.524 | -487 | - |
| BSA-AB | 3 | -1189 | -2.655 | 306 | 1.185 | -747 | -2.24 |
| 7 | -962 | -2.127 | 315 | 1.017 | -638 | -2.091 | |
| 10 | -831 | -1.051 | 1122 | 0.573 | -976 | -1.834 | |
| BSA-HAB | 3 | - | - | 920 | 0.81 | -460 | - |
| 7 | - | - | 1015 | 0.968 | -507 | - | |
| 10 | - | - | 1042 | 1.19 | -521 | - | |
| BSA-SOG | 3 | - | - | 113 | 0.377 | -56 | - |
| 7 | - | - | 214 | 0.514 | -107 | - | |
| 10 | - | - | 562 | 0.873 | -281 | - |
Table 2: CV for BSA with azo dyes (scan rate, 100 mV.s-1, concentration of BSA-2 × 10-5 M; dye concentration (× 10-4) -3, 7 and 10)
| Drug | Est. Free energy of binding (kcal/mol) | Est. Inhibition constant,(Ki.µM) |
vdW+ H-bond+dissolve energy (kcal/mol) |
Electrostatic energy (kcal/mol) | Total intermolecular energy (kcal/mol) |
Frequency | Intert. Surface |
|---|---|---|---|---|---|---|---|
| AB | -6.93 | 8.37 | -7.54 | +0.00 | -7.54 | 80% | 560.9 |
| HAB | -6.81 | 10.20 | -7.70 | -0.04 | -7.75 | 80% | 575.6 |
| SOG | -6.07 | 35.76 | -7.26 | -0.01 | -7.27 | 70% | 596.5 |
Table 3: Estimated free energy, inhibition constant, electrostatic energy and total intermolecular energy of the BSA with AB, HAB and SOG
Further, when more AB dye added to the BSA solution, the oxidation peak current (Ipa) is decreased and the reduction peak current (Ipc) is increased, whereas in HAB and SOG the Ipa increased. With the addition of adenine to the dyes, the voltammetric oxidation peak currents increased with positive shift, indicating that, there exist interactions between the dyes and BSA/ adenine. The drop of the voltammetric reduction current in the presence of BSA/adenine may be attributed to slow diffusion of the BSA/adenine binding with dyes. The current charge (whether an increase or decrease) is due to the interaction of the dyes with adenine or BSA. The results indicated that the binding reaction happened in the reaction solution and the electrode process was irreversible.
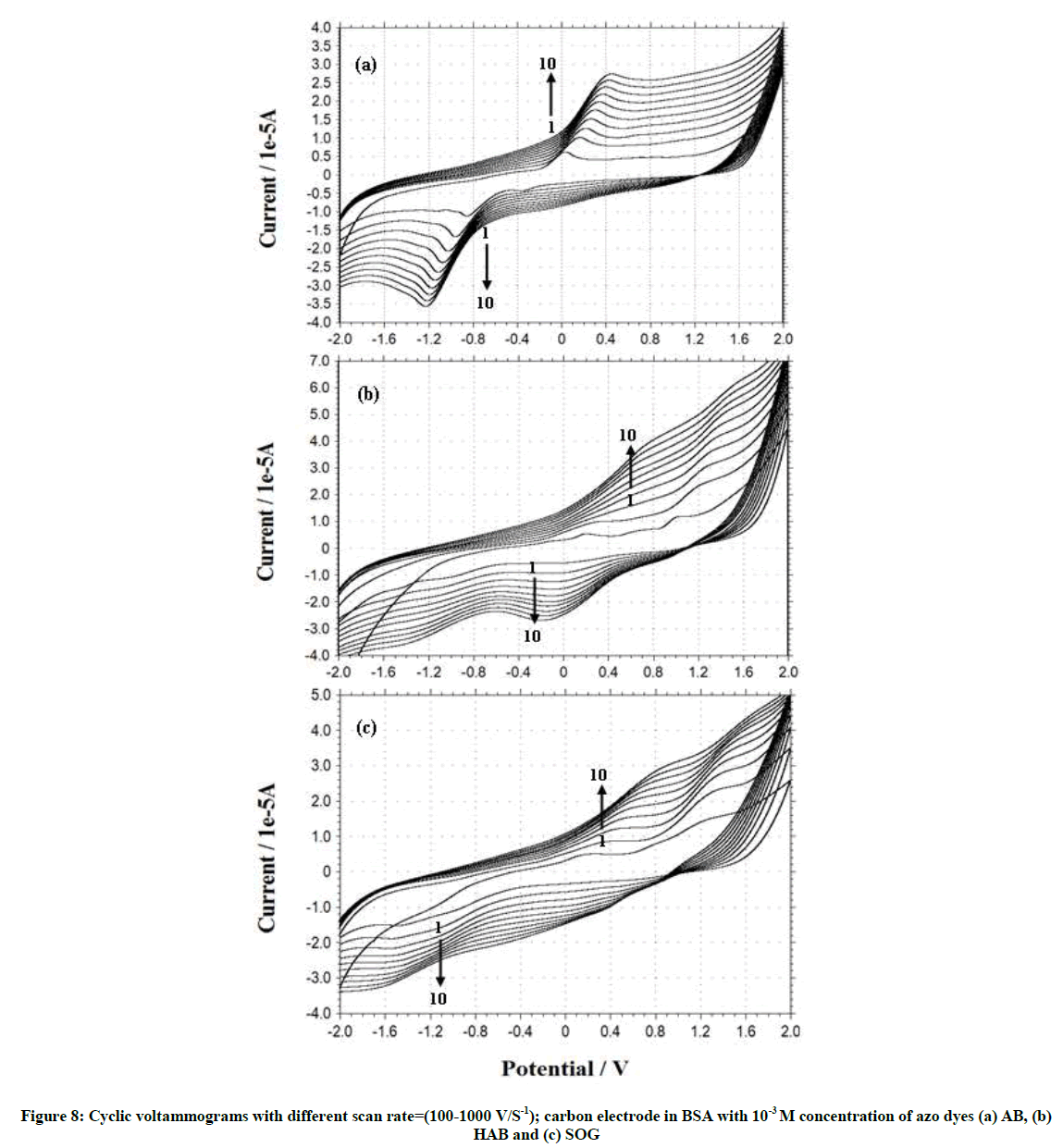
Effect of scan rate
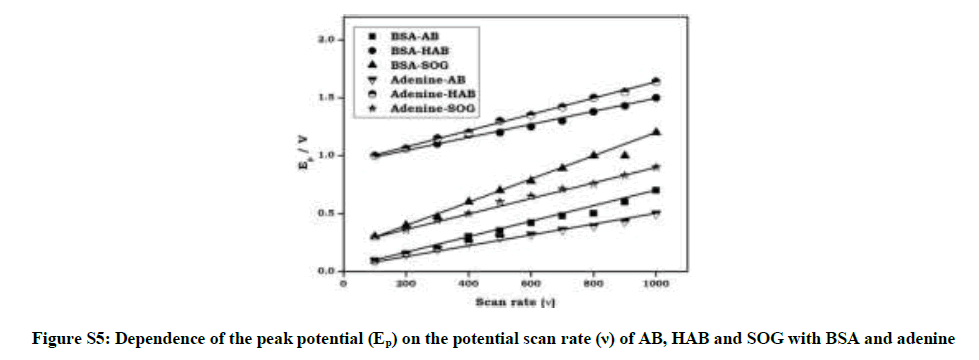
Cyclic voltammograms and the effect of scan rate on the peak current of azo dyes in BSA and adenine solution (pH~7.4) were investigated. From the cyclic voltammograms with scan rate (Figure 6), it can be seen that the electrode reaction of the dyes in the BSA and adenine are irreversible processes. The peak current was proportional to the root of scan rate over a range of 100 to 1000 mV.s-1.
The electrochemical reaction of dyes-BSA/adenine solution had the characteristics of the strong redox behavior of the reversible electrode process. So Laviron’s Equation [37,38] may be used to evaluate the kinetic constants of electrode reaction in the absence and presence of protein.
Ep = E0+RT/(αnF) [ln [(RTks)/ (αnF)] – lnν] (2)
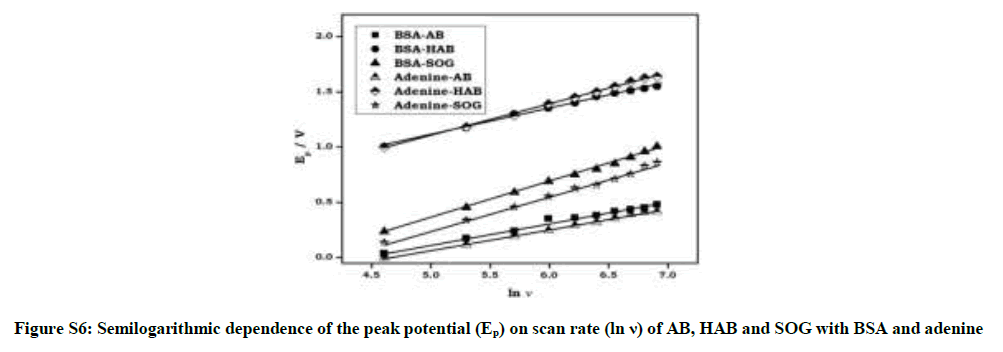
Where α is the electron transfer coefficient, ks the standard rate constant of the surface reaction, υ the scan rate, E0 the formal potential and n the electron transfer number. According to above equation, parameters of all the above dyes-BSA/adenine reaction solution were calculated with the same method and the results were also got from Figure S5 and Figure S6. Plot of Ep vs. υ is a well-defined straight line and αn value can be calculated from the slope and Ks from the intercept. Similarly, the plot of Ep vs. ln υ was shown in Figure S6, and was linear line. The E0 values of azo dyes can be determined from Figure S5 on the ordinate by extrapolating the line to υ=0.
Measurement of stoichiometry of BSA-dye complex
According to Li and Min, [39] the composition and the equilibrium constant can be analyzed derived from the changes of peak current (Ip). The electrochemical parameters of azo dyes-BSA or azo dyes-adenine reaction were calculated and compared to distinct the electrochemical activity of the formed bio-complex.
log[ΔI/(ΔImax-ΔI)] = logβs+m log[Dye] (3)
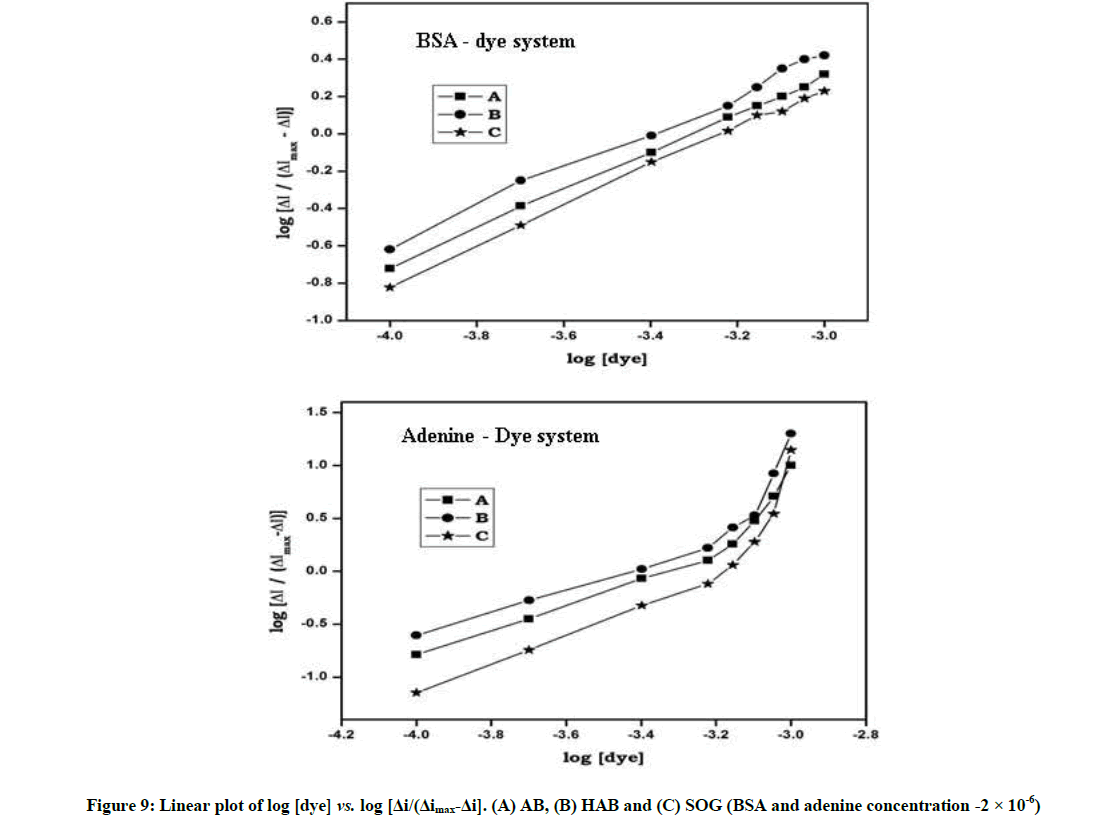
Where, ΔI was the peak current difference in the absence and presence of BSA, ΔImax corresponded to the obtained value when the concentration of the azo dye was much higher than that of BSA. CBSA, [BSA], [BSA-Dye] were corresponding to the total, free and bound concentration of protein in the solution, respectively. From Equation 3), the relation of log[ΔI/(ΔImax-ΔI)] with log[Dye] was calculated and plotted in Figure 7. From the intercept and slope m and βs were deduced, which indicated that the azo dyes binding to BSA/adenine formed the electro-inactive complex.
Further, the plot of log[Dye] vs. log [ΔI/(ΔImax-ΔI)] was linear in the BSA system (Figure 7) indicates that these dyes formed a single complex with protein, whereas in adenine system the plot of log[Dye] vs. log[ΔI/(ΔImax-ΔI)] was non-linear indicate that the complex is not a single complex between dyes and adenine.
Molecular docking studies
Molecular docking methods provide a deep-rooted approach to rank potential ligands (dyes) with respect to their ability to interact with a given target. The crystal structure of BSA was obtained from the Protein Data Bank (PDB code 4F5S) [24-26].
BSA biomolecule has three homologous domains (I, II, III), each one being composed of two subdomains (IA, IB, IIA, IIB, IIIA and IIIB). In BSA, subdomains IIA and IIIA are the favoured binding regions for aromatic and heterocyclic ligands. Therefore, we have searched these regions for favourable binding sites for dyes via molecular docking calculations. The best energy ranked result is shown in Table S4 and Figure 8. It can be seen that the azo dyes were bind into the BSA central cavity, which is in accordance with the conclusion of fluorescence experiment. It is found that a certain degree of distortion has happened on the azo dyes with BSA.
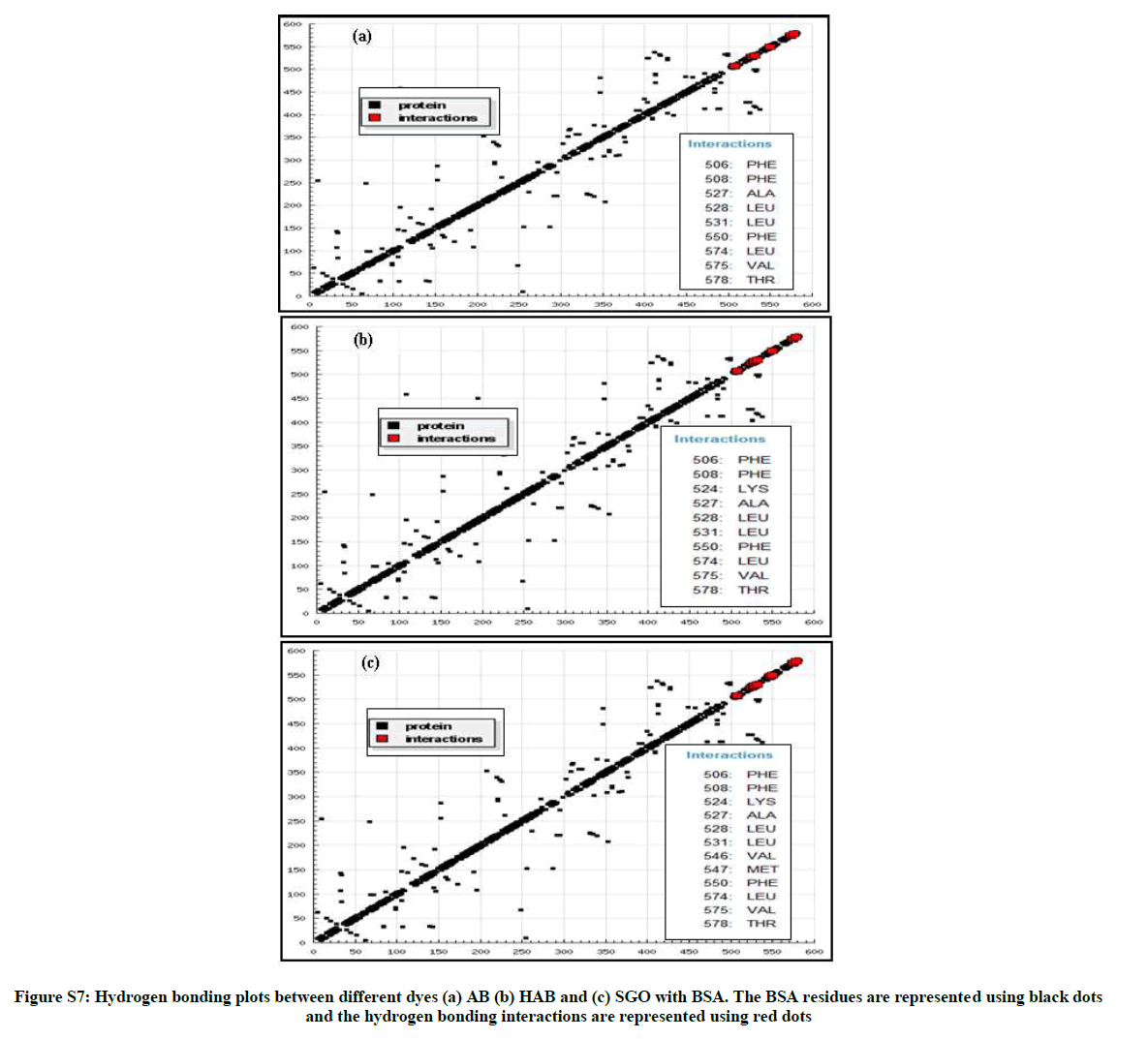
From the hydrogen bonding plot and the two-dimensional schematic representation (Figure S7 and Figure 9), it can be seen that the corresponding dyes are located within the binding pocket of subdomain IIIA and the AB molecule is surrounded by 9 amino acid residues (Leu- 387, PHE-507, PHE-508, ALA-527, LEU-528, LEU-531, PHE- 550, LEU-574, VAL575, THR-578) in the same way HAB and SOG molecules are surrounded by 10 and 12 amino acid residues.
It is important to note that the all the three corresponding azo dyes are adjacent to more hydrophobic residues, so hydrophobic forces may play major role in the binding. Further, hydrogen bonds not observed between the hydroxy groups of the azo dyes with amino acid residues of BSA which is in good agreement with the results of binding mode study.
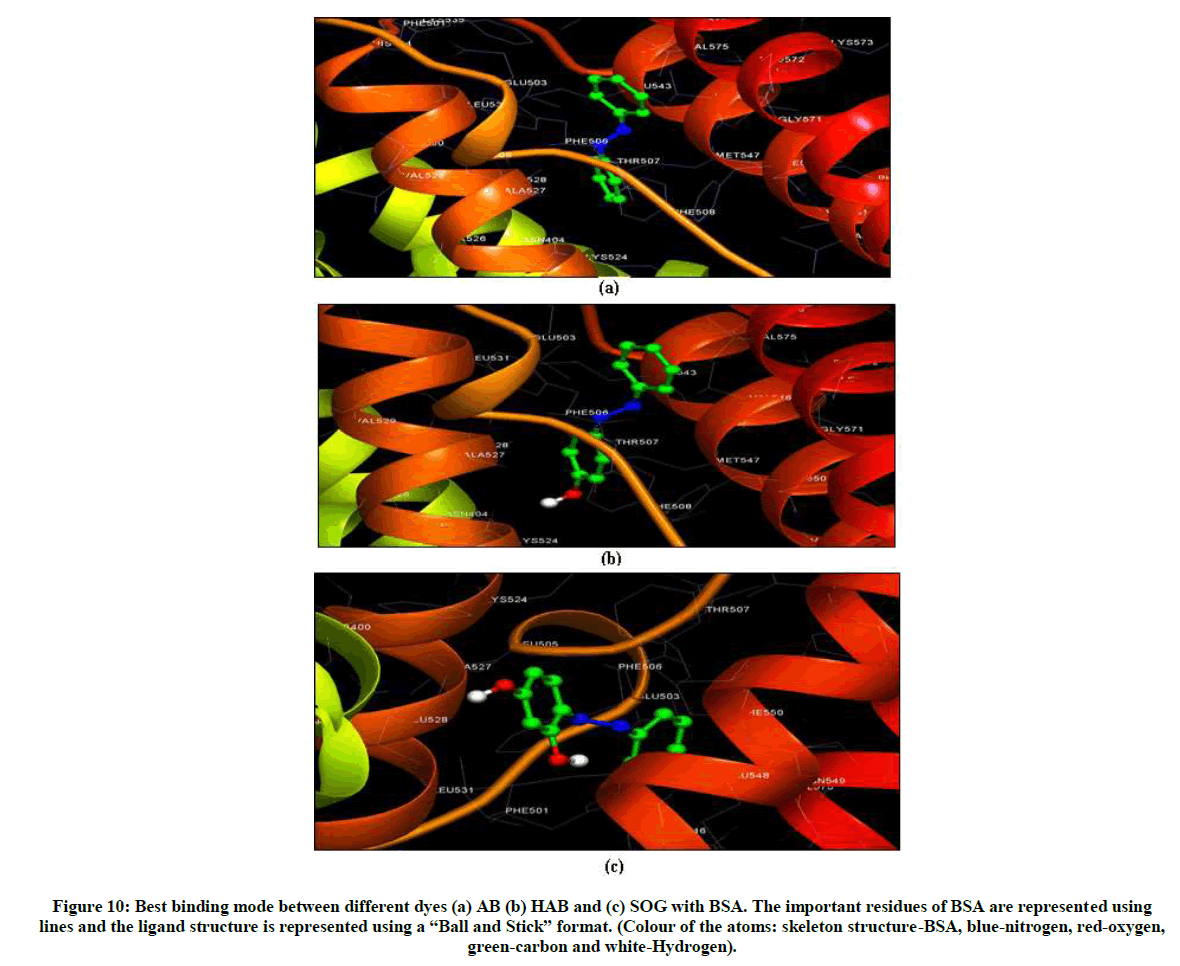
The list of interactions and its corresponding values are listed in Tables S2, S3 and S4 and shown Figure 10. The estimation free energy binding constant for the binding of AB, HAB and SOG to BSA were, 6.93, -6.81 and -6.03 kcal/mol respectively. These results are very secure to those obtained by the above mentioned experimental method. Combined with the spectral and electrochemical analysis, we believed that hydrophobic bonds, van der Waals interaction and electrostatic interactions are the driving forces in the binding of the dyes to BSA, rather than the hydrogen bonding interactions. Table S4 shows estimated free energy, van der Waals and hydrogen bonding interaction values of above discussed dyes with BSA. From this table, AB-BSA has more negative free energy than other two dyes indicate that AB molecule has more affinity to bind the active site of BSA than other dyes. As a consequence of this, the addition of a hydroxy group to the aromatic ring resulted in a decrease in the binding affinity [40].
| Hydrophobic | π-π | Other |
|---|---|---|
| C7 [3.65]–ALA527 (CB) | C4 [3.90]–PHE506 (CB) | N2 [3.69]–PHE506 CD2) |
| C8 [3.19]–ALA527 (CB) | C5 [3.78]–PHE506 (CB) | C2 [3.47]–THR578 CG2) |
| C9 [3.65]–ALA527 (CB) | C1 [3.80]–PHE506 (CD2) | C3 [3.45]–THR578 CG2) |
| C11 [3.45]–LEU528 (CB, D1) | C2 [3.68]–PHE506 (CD2, E2) | |
| C2 [3.50]–LEU531 (CD1) | C8 [3.86]–PHE508 (CZ) | |
| C12 [3.76]–LEU531 (CD2) | C9 [3.41]–PHE550 (CD2, E2, CG, CZ) | |
| C6 [3.52]–LEU574 (CD2) | C8 [3.72]–PHE550 (CD1, CE1, CZ) | |
| C4 [3.54]–VAL575 (CB, CG2) | ||
| C5 [3.33]–VAL575 (CG2) |
The strong interactions are mentioned in bold letters
Table S2: List of interactions between BSA and AB
| π-π | Other | |
|---|---|---|
| C1 [3.73]–ALA527 (CB) | C11[3.80]–PHE506 (CB) | N1 [3.35]–PHE506 (CD2, CE2) |
| C2 [3.41]–ALA527 (CB) | ||
| C3 [3.68]–ALA527 (CB) | C7 [3.73]–PHE506 (CD2) | H10 [3.31]–LYS524 (CB) |
| C5 [3.49]–LEU528 (CB, CD1) | C8 [3.77]–PHE506 (CD2) | |
| H10 [3.69]–LEU528 (CB) | ||
| C8 [3.35]–LEU531 (CD1) | C2 [3.78]–PHE508 (CZ) | C8 [3.33]–THR578 (CG2) |
| C2 [3.60]–PHE550 (CD1, CE1, CG) | C9 [3.40]–THR578 (CG2) | |
| C6 [3.48]–LEU531 (CD2) | ||
| C12 [3.83]–LEU574 (CD2) | C3[3.50]–PHE550 (CD1, CD2, CE1, E2) | |
| C10 [3.66]–VAL575 (CB, CG1, CG2) | ||
| C11 [3.10]–VAL575 (CB, CG2) |
Table S3: List of interactions between BSA and HAB
| Hydrophobic | π-π | Other |
|---|---|---|
| C2 [3.69]–ALA527 (CB) | C9 [3.52]–PHE506 (CB) | C3 [3.47]–LYS524 (CB, CG) |
| C5 [3.81]–LEU528 (CD1) | C10 [3.62]–PHE506 (CB) | O2 [3.50]–LEU531 (CD2) |
| C5 [3.69]–MET547 (CG) | C8 [3.36]–PHE508 (CE2) | H10 [3.38]–LEU531 (CD2) |
| C7 [3.83]–LEU574 (CD2) | C9 [3.72]–PHE508 (CE2) | O2 [3.33]–VAL546 (CG1) |
| C8 [3.34]–LEU574 (CD2) | C1 [3.87]–PHE550 (CG) | H10 [2.90]–VAL546 (CG1) |
| C9 [3.54]–LEU574 (CD2) | C2 [3.50]–PHE550 (CD2, E2, CG, CZ) | H9 [3.83]–MET547 (CE) |
| C10 [3.79]–VAL575 (CG2) | C3 [3.46]–PHE550 (CD2, CE2) | N2 [3.40]–PHE550 (CB,CD1, CG) |
| C11 [3.49]–THR578 (CG2,OG1) | ||
| C12 [3.84]–THR578 (CG2) |
Table S4. List of interactions between BSA and SOG
Figure 10: Best binding mode between different dyes (a) AB (b) HAB and (c) SOG with BSA. The important residues of BSA are represented using lines and the ligand structure is represented using a “Ball and Stick” format. (Colour of the atoms: skeleton structure-BSA, blue-nitrogen, red-oxygen, green-carbon and white-Hydrogen).
Conclusion
Interactions between BSA and adenine with three azo benzenes have been investigated by using different biophysical techniques. BSA fluorescence can be statically quenched by all the dyes, which implies that the corresponding dyes can bind to BSA and adenine to form like protein-ligand complex. Azo dyes can bind to BSA with stoichiometric ratio of 1:1 and the protein – ligand complexes are stabilized mainly by hydrophobic and van der Waals interaction. AB molecule has more E value than other two dyes in BSA and adenine systems which mean the possibility of energy transfer from AB molecule to BSA is higher than HAB and SOG molecule. IR spectral analysis shows, the azo group (N=N) stretching frequency for AB, HAB and SOG were shifted in the BSA complex and the changes in spectral shape and band position of pure BSA, AB, HAB and SOG in the corresponding interactions confirm the formation of the protein-dye complexes. The concentration of the dyes added to the BSA solution, a reductive peak appeared in AB, but this peak is not noticed in HAB and SOG. When more concentration of AB dye added to the BSA solution, the oxidation peak current (Ipa) decreased and reduction peak current (Ipc) increased, where as Ipa increased in HAB and SOG. Molecular docking studies specifies that the corresponding dyes are located within the binding pocket of sub-domain IIIA in BSA and the hydrophobic forces were plays a major role in the binding. Compared to HAB and SOG, AB molecules contribute substantially higher binding efficiency with BSA/adenine. As a consequence of this, the addition of a hydroxy group to the aromatic ring resulted in a decrease in the binding affinity.
Acknowledgement
This work was supported by the Council of Scientific Industrial Research [No. 01(2549)/12/ EMR-II], New Delhi, India and the University Grants Commission [F. No. 41-351/2012 (SR)], New Delhi, India.
References
- B.P. Kamat, J. Seetharamappa, Pol. J. Chem., 2004, 78, 723-732.
- K. Golka, S. Kopps, Z.W. Myslak, Toxicol. Lett., 2004, 151, 203-210.
- N.A.G.A. Refat, Z.S. Ibrahim, G.G. Moustafa, K.Q. Sakamoto, M. Ishizuka, S.J. Fujita, Biochem. Mol. Toxicol., 2008, 22, 77-84.
- R. Rebane, I. Leito, S. Yurchenko, K. Herodes, J. Chromatogr. A., 2010, 1217, 2747-2757.
- G. Kesiunaite, A. Linkeviciute, E. Naujalis, A. Padarauskas, Chromatographia., 2009, 70 1691-1695.
- L. Li, H.W. Gao, J.R. Ren, L. Chen, Y.C. Li, J.-F. Zhao, H.P. Zhao, Y. Yuan, BMC Struct. Biol., 2007, 7, 16-26.
- R.H. Wesley, Coord. Chem. Rev., 1996, 149, 347-365.
- T.O. Hushcha, A.L. Luik, Y.N. Naboka, Talanta., 2000, 53, 29-34.
- D.C. Carter, J.X. Ho, Adv. Protein Chem., 1994, 45, 153-203.
- T. Peters, All About Albumin, Biochemistry, Genetics and Medical Applications, Academic Press, San Diego, 1996.
- X.M. He, D.C. Carter, Nature., 1992, 358, 209-215.
- T. Peters, Adv. Protein Chem., 1985, 37, 161-245.
- T. Jianniao, L. Jiaquin, H. Zhide, C. Xingguo, Bioorg. Med. Chem., 2005, 13, 4124-4129.
- D. Silva, C.M. Cortez, S.R.W. Louro, Spectrochim. Acta Part A., 2004, 60, 1215-1223.
- J. Prema kumari, A. Antony muthu prabhu, G. Venkatesh, V.K. Subramanian, N. Rajendiran, J. Solu. Chem., 2011, 40, 327-347.
- G. Venkatesh, J. Saravanan, N. Rajendiran, Bull. Chem. Soc. Japan., 2014, 87, 283-293.
- A. Antony muthu prabhu, G. Venkatesh, R.K. Sankaranarayanan, N. Rajendiran, Indian J. Chem., 2010, 49A, 407-417.
- A. Antony muthu prabhu, G. Venkatesh, N. Rajendiran, J. Fluoresc., 2010, 20, 961-972.
- N. Rajendiran, R.K. Sankaranarayanan, Carbohydr. Polym., 2014, 106, 422-431.
- N. Rajendiran, G. Venkatesh, J. Saravanan, J. Mol. Struc., 2014, 1072, 242-252.
- Z. Bikadi, E. Hazai, J. Chem. Inf., 2009, 1, 1-15.
- T.A. Halgren, J. Comput. Chem., 1998, 17, 490-519.
- G.M. Morris, D.S. Goodsell, J. Comput. Chem., 1998, 19, 1639-1662.
- N. Rajendiran, J. Thulasidhasan, Spectrochim. Acta A., 2015, 144, 183-191.
- N. Rajendiran, J. Thulasidhasan, Canadian Chemical Transactions., 2015, 3, 291-307.
- N. Rajendiran, J. Thulasidhasan, Int. Lett. Chem. Phy. Ast., 2015, 59, 170-187.
- S.Y. Bi, L.L. Yan, B.B. Wang, J. Lumin., 2011, 131, 866-873.
- L.L. He, X. Wang, B. Liu, J. Solution. Chem., 2010, 39, 654-664.
- J.R. Lackowicz, Principles of Fluorescence Spectroscopy, 3rd edi., Plenum Press, New York, 2006.
- J. Thipperudrappa, D.S. Biradar, M.T. Lagare, S.M. Hanagodimath, S.R. Inamdar, J.S. Kadadevaramath, J. Photochem. Photobio A: Chem., 2006, 177, 89-93.
- S.M.T. Shaikh, J. Seetharamappa, P.B. Kandagal, D.H. Manjunatha, S. Ashoka, Dyes Pigm., 2007, 74, 665-670.
- O.K. Abou-Zied, O.I.K. Al-Shihi, J. Am. Chem. Soc., 2008, 130, 10793-10801.
- Y.Z. Zhang, B. Zhou, X.P. Zhang, P. Huang, C.H. Li, Y. Liu, J. Hazard. Mater., 2009, 163, 1345-1352.
- A. Papadopoulou, R.J. Green, R.A. Frazier, J. Agric. Food Chem., 2005, 53, 158-163.
- B. Valeur, Principles and Applications, Wiley, New York, 247-272.
- J. R. Lakowicz, G. Weber, Biochem., 1973, 12, 4161-4170.
- E. Laviron, J. Electronanal. Chem, 1974, 52, 355-393.
- E. Laviron, J. Electroanal. Chem., 1979, 101, 19-28.
- N. Q. Li, J. Min, Chin. J. Anal. Chem., 1989, 17, 346-354.
- C. Dufour, O. Dangles, Biochim. Biophys. Acta., 2005, 1721, 164-173.