Research Article - Der Pharma Chemica ( 2018) Volume 10, Issue 10
Molecular Identification of Pectinolytic Streptomyces sp. GHB5 and Optimization of Polygalacturonase Production
Nadia H Abd El-Nasser1, El Sayed M El-Mahdy2, Wafaa Gh Shosha2 and Gehad H El Sayed1*
1Department of Microbial Chemistry, National Research centre, Dokki, Giza, Egypt
2Department of Chemistry, Facualty of Science, Helwan University, Egypt
- *Corresponding Author:
- Gehad H El Sayed
Department of Microbial Chemistry
National Research centre
Dokki, Giza, Egypt
Abstract
Streptomyces sp. GHB5 was isolated from Egyptian soil, optimized for polygalacturonase production and identified as Streptomyces sp. GHB5 by 16S rRNA sequencing and submitted to gene bank data base with accession number LC405930. Submerged fermentation was carried out using Streptomyces sp. GHB5 for polygalacturonase production. Different physico-chemical and nutritional factors were optimized for maximum polygalacturonase production. Results revealed that the maximum polygalacturonase production come to 34.09 U/mg specific activity. The best obtained conditions were achieved using broth medium with working volume of 20 % (v/v), containing 1.5% (w/v) lemon peel and 0.28% (w/v) potassium nitrate with initial medium pH of 7.5, when incubated at 30°C and 180 rpm for 6 days with an inoculum size of 4% (v/v) of 5 days inoculum age.
Keywords
16S rRNA sequencing, Streptomyces sp. GHB5, Polygalacturonase production, Optimization.
Introduction
Biotechnology processes for enzyme creation is no longer an academic, it is probably useful alternative proposition for the future [1,2]. Enzymes are natural biocatalysts. They are produced by living organisms to raise the rate of the immense and diverse group of chemical reactions needed for life [2].
Pectinase is a general term for enzymes, such as pectolyase, pectozyme and polygalacturonase [3,4] which responsible for pectin destruction from different types of plants [4,5]. By 2011, the world industrial market for pectinases had revenues of 70 million dollars, [6] accounting for at least 17% of a 1200 million dollars share of the food and beverages enzymes, a segment which in turn reached 1600 million dollars by 2016 [7]. The top stake pectinases hold in the enzyme marketing area is related to its large-scaled application, spanning any process that requires pectin breakdown [8]. Pectin is one of the most widely present polysaccharide in nature. Pectin is mainly found in large number of plant tissues such as lemon peel, orange peel, pulp, sugar beet, wheat bran, rice bran, and other citruses [9].
Annually, about 34 million tons of citrus wastes (peel, seeds) are produced from the processing of citrus fruits [10-13]. Although a part of these residues is used as a part of the diet of monogastric animals, its accumulation in the biosphere causes several environmental problems. Also, citrus peel is usually an important and economical raw material to produce enzymes, mainly pectinase, by fermentative techniques. This material contains almost all nutrients needed for microorganism growth, and several researches have been studied for the production of many important metabolites including enzymes [13,14]. To reduce the cost of production, lemon peel (pectic waste) as a substrate is used rather than synthetic cellulose owing to their reasonable fair cost, high enzyme production capacity etc. The decrease in cost causes an economically cheap easy method for pectinase production. It is an important issue to handle the residue, both for the comprehensive utilization of pectic resources and for the prevention of polluting the environment [13].
The capability to synthesize pectinolytic enzymes is very common in categories of microorganisms [15]. Polygalacturonase are obtained from large number of fungi, bacteria, actinomycetes and also by higher plants [4,16-19]. Pectinase production by microbial growing on complex substrates, as fruit residues, is a complicated process and controlled by several factors, so the identification of optimal growth conditions for the highest pectinase productivity is not fairly easy task [20,21]. Pectinase enzymes are produced by three approaches including solid-state fermentation, submerged fermentation, and surface culture. Solid-state and submerged fermentation systems are preferred for pectinase production by microorganisms because it can be performed using agricultural and industrial wastes, which are cheap carbon sources. At the same time these systems have more efficiency and fewer waste products. Pectinase production is affected by culture medium compositions, especially carbon and nitrogen sources, and different physical or chemical parameters such as pH, temperature, and aeration [9,22,23].
Actinomycetes usually defined as Gram-positive bacteria that contain high G+C (>55%) content in their DNA [24,25]. It is known by forming branching filaments with hyphae and asexual spores. In addition, they were considered an intermediate group between bacteria and fungi but now are recognized as prokaryotic micro-organisms [25-27]. Most soil actinomycetes are neutrophils, growing between pH 5.0 and 9.0 with an optimum near to neutrality. On top of that, most soil actinomycetes are strict aerobes [25,27]. Also, they are really important in soil biodegradation and humus formation [28], and produce several volatile substances like geosmin that responsible for the wet earth characteristic odor [25,29].
In 16S rRNA tree, actinomycetes form a definite phylogenetic line and have been a major technological importance in the past years, with the discovery of numerous number of metabolites produced by its various genera [30].
The present investigation was carried out for molecular identification of Streptomyces sp GHB5 and to improve polygalacturonase production from this strain.
Materials and Methods
Molecular Identification Streptomyces sp GHB5 Isolate
DNA extraction
Streptomyces sp. GHB5 was grown on starch nitrate agar plates where a single colony of it picked up and streaked on starch nitrate agar plates and incubated at 28ºC for 5 days for obtaining heavy growth to be used for DNA extraction. DNA was extracted with GeneJet genomic DNA purification Kit (Thermo K0721).
PCR amplification
Amplification was as follows: Initial denaturation at 95°C for 10 min followed by 35 cycles of denaturation at 95°C for 30 s, annealing at 65°C for 1 min, and Extension at 72°C for 1 min 30 s and final Extension at 72°C for 10 min. PCR Purification Kit (Thermo K0701) was finally used for PCR product purifications.
Purification of the PCR product
PCR products were purified using PCR cleanup to the PCR product using GeneJET™ PCR Purification Kit (Thermo K0701) according to the supplierʹs instructions.
Sequencing of the amplified PCR product
Sequencing to the PCR product on GATC Company by use ABI 3730xl DNA sequencer by using the set of primers 27F (5′-AGA GTT TGA TCC TGG CTC AG-3′) and 1294R (5′-GGT TAC CTT GTT ACG ACTT-3′), Data were submitted to GenBank database.
Sequence similarities and phylogenetic analysis
The nucleotide sequence was compared to the GenBank database in the national Center for Biotechnology Information using the BLAST program (https://www.ncbi.nlm.nih.gov/) [31]. Selected sequences of the greatest similarity to the 16S rRNA sequence of the Streptomyces sp. GHB5 isolate were aligned and generating the phylogenetic tree. The partial 16S rRNA sequence of Streptomyces sp. GHB5 was assigned in the GenBank with accession no. LC405930.
Strain
Streptomyces sp. GHB5 was isolated from Egyptian soil and screened for polygalacturonase production following a previous research [32].
Fermentation substrate
lemon peels (Lp) was obtained from local market in Egypt, cut into small pieces (~2 mm), washed with tap water several times in order to remove all water soluble compound then dried and grinded.
Fermentation medium
Lemon peel nitrate medium [32] used for enzyme production, containing (g/l): lemon peel, 20 g, KNO3 2.0 g, K2HPO4 (anhydrous), 1.0 g, MgSO4. 7H2O, 0.5 g, NaCl, 0.5 g, CaCO3 3.0 g, FeSO4. 7H2O, 0.01 g, Distilled water 1000 ml. was utilized for the inoculum medium and as cultivation medium. The medium was sterilized by autoclaving at 121ºC for 20 min.
Polygalacturonase assay
Polygalacturonase (P.G) was determined according to [33] as described in previous research [32]. One unit (1U) of enzyme activity is equal to the 1 μMol of reducing sugars released per ml per min, measured in terms of D-galacturonic acid, produced as a result of enzyme substrate reaction.
Protein determination
The protein concentration was determined in the enzyme solution according to [34]. Bovine serum albumin was used as a standard.
Optimization of polygalacturonase production different
Seed inoculum age
Seed inoculum of Streptomyces sp. was prepared by growing it on starch nitrate agar slants at 28ºC for 1-10 days, then the slant cultures washed with 5 ml of sterile distilled water to prepare inocula. Flasks were loaded with lemon peel nitrate medium, at 20% v/v of different seed inoculum age and allowed to grow at 28ºC for 5 days on a rotary shaker at 180 rpm.
Fermentation time per day
Streptomyces sp. with the best inoculum age was cultivated in 250 ml Erlenmeyer flasks containing 20 % v/v of production medium. Flasks incubated for 1, 2, 3, 4, 5, 6, 7, 8, 9 and 10 days under shaking (180 rpm) conditions at 28ºC.
The effect of inoculum size
250 ml Erlenmeyer flasks containing 20 % v/v of production medium were inoculated with different volumes of spore suspension Streptomyces sp such as 1, 2, 3, 4, 5, 6, 7, 8, 9 and 10% (v/v) containing 4*107 spores/ ml (OD550=0.052). Flasks allowed to grow at 28ºC for the optimum incubation period on a rotary shaker at 180 rpm.
Initial pH of the medium
Streptomyces sp. was cultivated in 250 ml Erlenmeyer flasks containing 20 % v/v of production medium. The initial pH of the production medium was adjusted with 0.1 M NaOH or 0.1 M HCl from pH 4.0 to 9.0. Flasks incubated for the optimum incubation period under shaking (180 rpm) conditions at 28ºC.
Working volume
Sterptomyces isolate was cultivated in 250 ml Erlenmeyer flasks containing 10%, 20%, 40% and 60% (v/v) of production medium. Cultivated flasks were incubated for the best incubation period, at the best pH under shaking (180 rpm) conditions at 28ºC.
Incubation temperature
The best working volume of production medium was cultivated with Sterptomyces strain in 250 ml conical flasks. Flasks were incubated for the best incubation period, at the best pH, at 180 rpm and at different incubation temperatures (20ºC, 25ºC, 28ºC, 30ºC, 35ºC, 40ºC and 45ºC).
Shaker speed (rpm)
The best working volume of production medium was cultivated with Sterptomyces sp. in 250 ml conical flasks. Flasks then incubated for the optimum incubation period and optimum pH at optimum temperature at different rotation per min (rpm) (50, 100, 150, 180, 200 and 250 rpm).
Carbon sources
An equimolar carbon concentration of sucrose, fractose, D-manitol, arabinose, raffinose, lactose, starch and glycerol was added instead of lemon peel in conical flasks containing the best working volume of the basal medium with the same final concentrations of C and N of the lemon peel nitrate medium. Flasks were cultivated with Sterptomyces sp. at the optimum pH, then they were incubated at the optimum incubation period, at the optimum rpm and the optimum temperature.
Different concentrations of carbon source
Different concentrations of the optimum carbon source (0.5, 0.10, 0.15, 0.20, 0.25, and 0.3 w/v) have been added to the optimum working volume of production medium in 250 ml conical flasks at the optimum pH. Flasks were then incubated at the optimum incubation period at the optimum shaker speed and temperature.
Nitrogen sources
The optional concentration of potassium nitrate in the basal medium was replaced in equimolar by ammonium nitrate, peptone, urea, ammonium chloride and ammonium hydrogen phosphate, casein and peptone. The replaced nitrogen sources have been added to the optimum working volume of production medium in 250 ml conical flasks at the optimum pH with the optimum concentration of carbon source. Flasks were then incubated for the optimum incubation period at the optimum shaker speed and temperature.
Different concentrations of nitrogen source
Different concentrations of nitrogen source (0.1, 0.14, 0.18, 0.2, 0.24, 0.28, 0.3, 0.4 w/v) have been used. The optimum working volume of production medium was added to different concentration of nitrogen sources in 250 ml conical flasks at the optimum pH with the optimum concentration of carbon source. Flasks were then incubated at the optimum temperature and rpm for the optimum incubation period.
Results and Discussions
Molecular characterization of the pectinolytic Streptomyces sp. GHB5
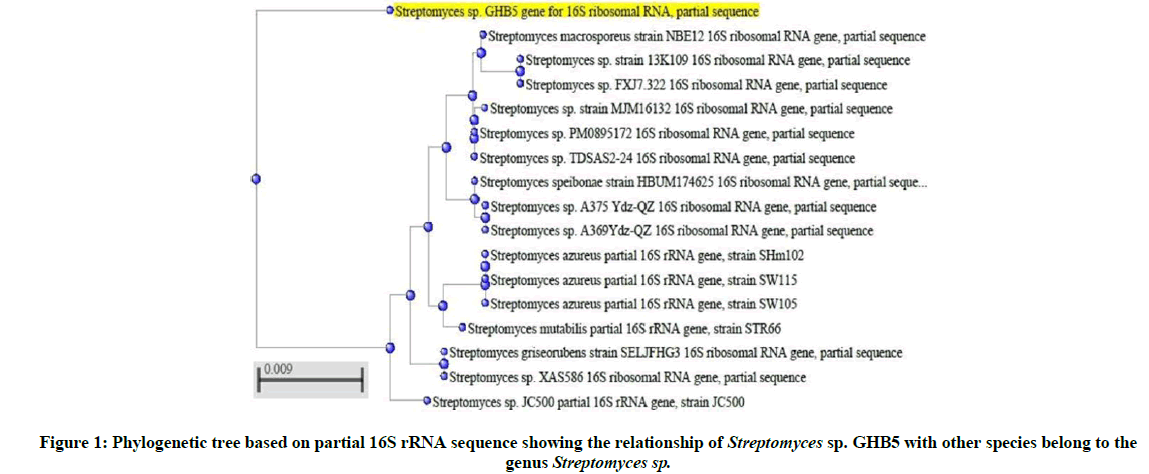
The 16S rRNA sequence of the Streptomyces sp. GHB5 was obtained and submitted to GenBank database. This submitted nucleotide sequence was provided a GenBank accession number LC405930. Based on the results of BLAST search using the sequence data the Streptomyces sp. isolate was found to be strongly related to Streptomyces sp. The phylogenetic tree was constructed for partial 16S rRNA sequences of the isolate and the closest matches from BLAST searches (Figure 1).
Optimization of polygalacturonase production
Physical factors are important in any fermentation for optimization of biochemical production. The key physical factors that determine the bioprocess are pH level, temperature, aeration and shaking speed [35].
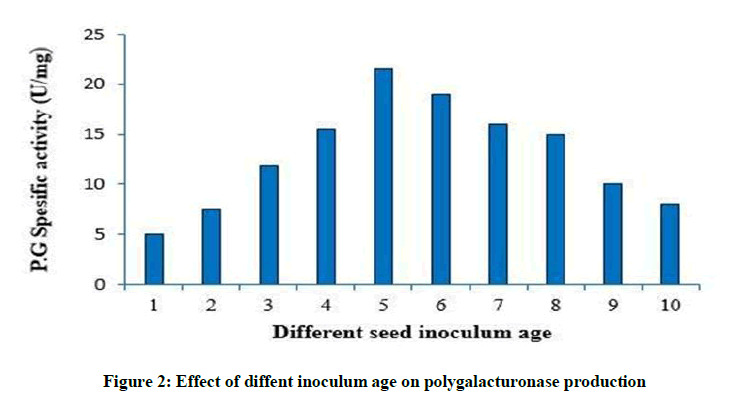
Effect of diffent inoculum age on polygalacturonase production
As shown in Figure 2, rapid increase in enzyme production was observed till the inoculum age of 5th day. Further increase in inoculum age led to reduction in enzyme activity.
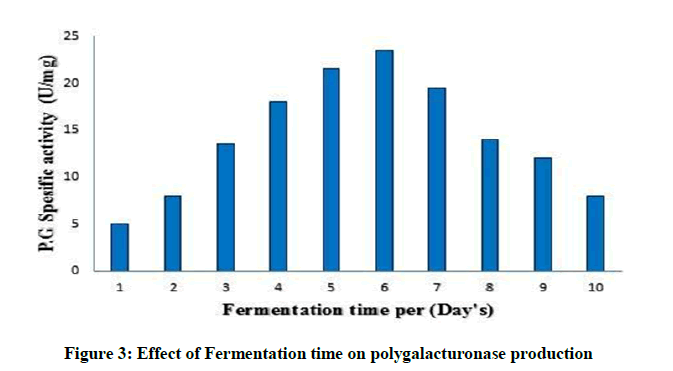
Effect of Fermentation time on polygalacturonase production
As shown in Figure 3, the level of enzyme production varies with the time duration of the fermentation process [36]. Maximum enzyme yield obtained at 6 days. Data showed that the polygalacturonase activity was decreasing after 6 days of the fermentation process. Further than this period the enzyme synthesis drastically reduced, probably owing to the depletion of essential nutrients in the medium and/or accumulation of harmful secondary metabolites or the cells may reach the decline phase and exhibited low polygalacturonase synthesis. This result is in agrement with similar observations of [37].
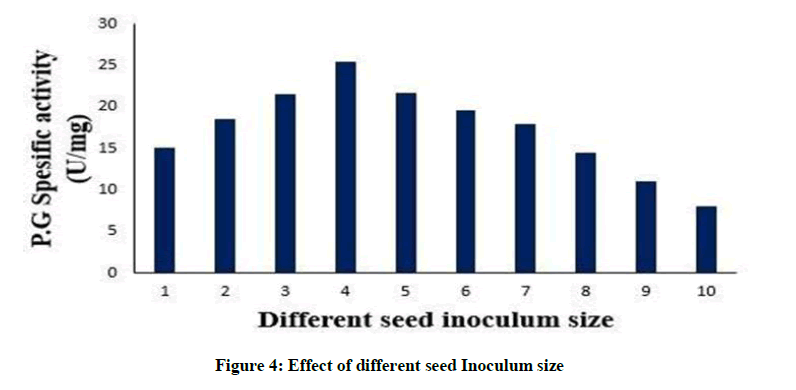
Effect of different seed Inoculum size
The initial load of microbes also influences the last amount of the enzyme synthesized. 4% (v/v) inoculum resulted in highest polygalacturonase production. With a rise in the inoculum percentage the enzyme production steadily increased, probably due to effective use of the substrate by the actively growing microbial cells, as depicted in Figure 4. A reduction in the enzyme production was noticed when 10% (v/v) inoculum was used. This might be owing to rapid depletion of nutrients and oxygen stress resulting from a high microbial load [2]. An enzyme activity of 136 U/ml was obtained from Bacillus circulans at an inoculum size of 5% [38]. While optimum pectinase production of 5.92 IU was reported with inoculum size of 10.0% (V/V) [2].
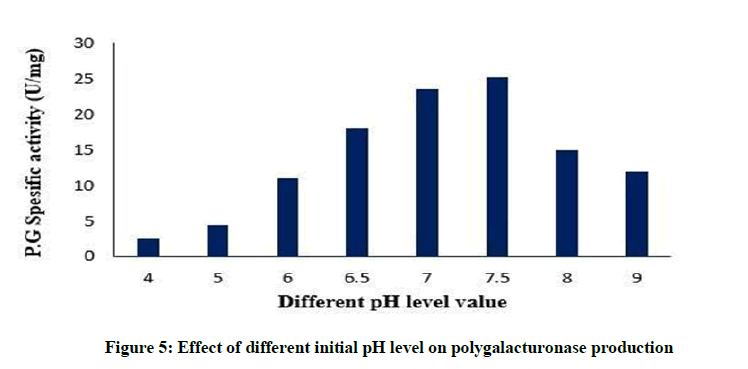
Effect of different initial pH on polygalacturonase production
pH level of the fermentation medium plays an essential role in identifying the level of metabolite synthesis. The stability of the microbial metabolite is also influenced by the hydrogen ion concentration of the medium [39]. In the present study, a medium pH of 7.5 supported maximum polygalacturonase production, as stated in Figure 5. This result is similar to [40]. It is mentioned that at high pH, the metabolic action of micro-organism may be suppressed and therefore it inhibits the enzyme production [41].
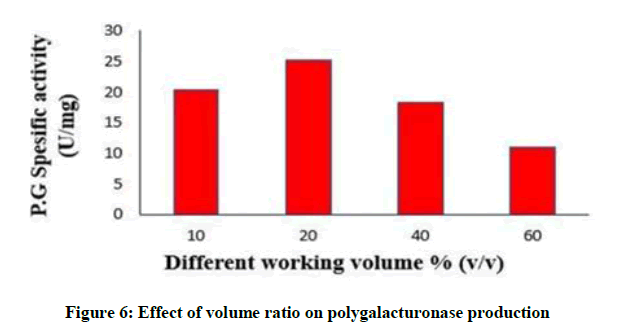
Effect of working volume on polygalacturonase production
Polygalacturonase production was affected by changes in volume ratio as described in Figure 6. The highest production was attained at a volume ratio of 20 % (v/v) yielded (25.3) U/mg of specific polugalacturonase activity. Enzyme activity lowered by increasing volume this may be a result of oxygen limit.
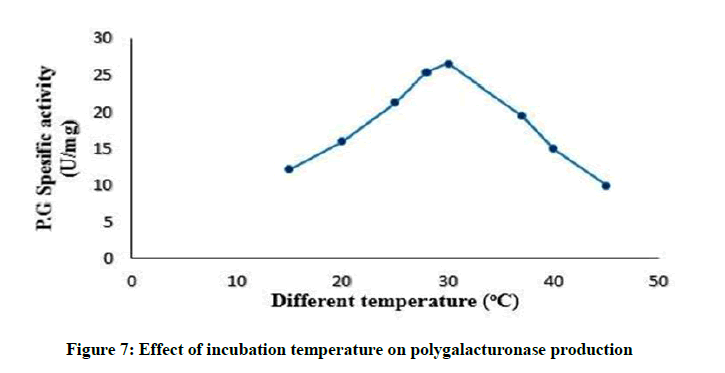
Effect of incubation temperature on polygalacturonase production
The incubation temperature greatly influences the microbial growth rate, enzyme secretion, enzyme inhibition and protein denaturation [36]. The best temperature for maximum enzyme production was 30°C, as presented in Figure 7. This results in agreement to [2,37,42,43]. Due to the mesophilic character of the Streptomyces isolate, this temperature may be responsible for better membrane permeability and therefore maximum polygalacturonase secretion into the medium. By increasing temperature above 30°C, the enzyme production lowered as a result of growth reduction and enzyme inactivation or suppression of cell viability [44]. At higher temperatures, damage occur to microorganisms by denaturing enzymes, transport carriers and other proteins thus decreasing enzyme production [2,45]. In contrast, low temperature values membranes solidify and may reduce the metabolism of the microorganism and consequently, the enzyme synthesis. This result is similar to observations of [37,44,46].
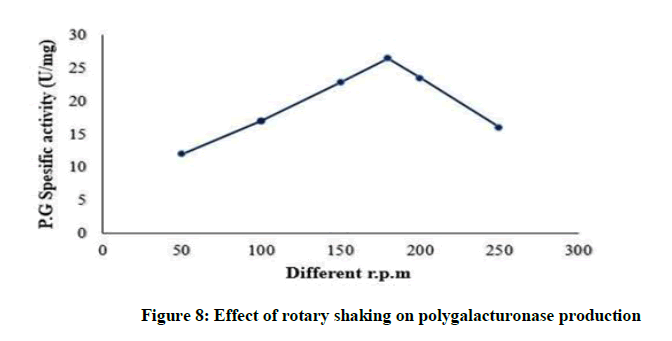
Effect of rotary shaking polygalacturonase production
By increasing agitation polygalacturonase production increases till reach the maximum yield at 180 rpm (26.5 U/mg) as shown Figure 8, then polygalacturonase production reduced by increase in rotary shaking per mint. It was stated that enzyme activity increased together with increments in agitation rates up to a threshold level from which shear stress caused damage to cells and resulted in poor biomass development and metabolite synthesis [8].
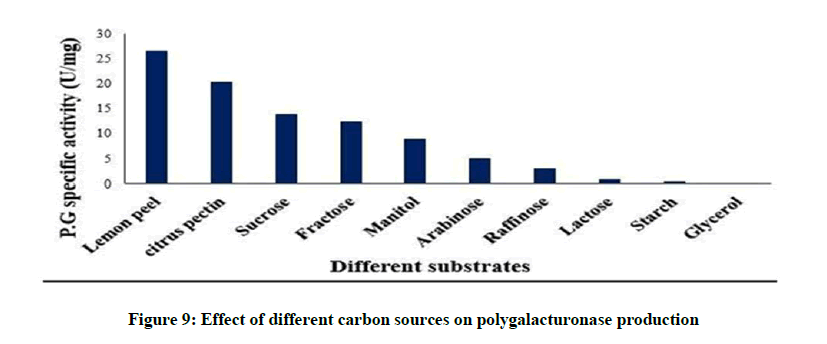
Effect of different carbon sources on polygalacturonase production
The effect of different carbon sources on Polygalacturonase production is shown in Figure 9. Lemon peel was the most effective carbon source for Polygalacturonase production (26.5 U/mg). In our present study, the effect of sucrose was more than the other carbon sources tested. Fructose was the second best supplementary carbon source. Starch gave the lowest enzyme activity. It was reported that different carbon sources have varied effect on the production of extracellular enzymes especially pectinase strains [47,48].
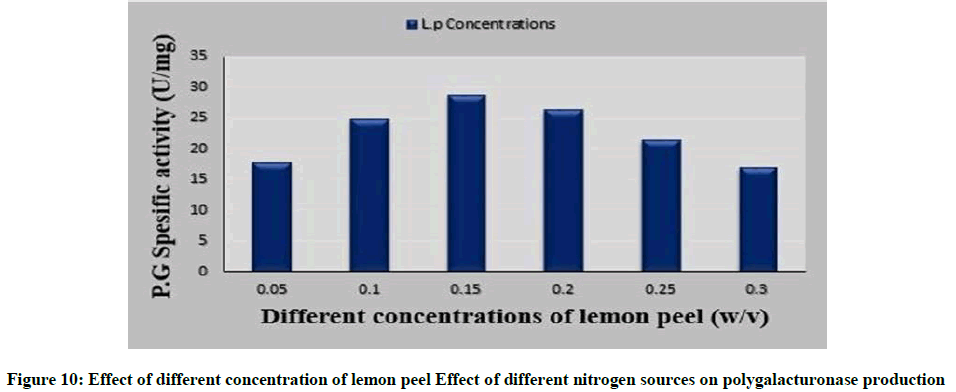
Effect of different concentration of lemon peel (the best carbon source) on polygalacturonase production
Availability of good amount of an utilizable substrate in the fermentation medium supports high production of the enzyme. Figure 10 showed that by increasing the concentration of lemon peel powder more than 1.5%, the polygalacturonase production was noticed to lower. This can be linked to phenomenon of catabolite repression, where galacturonic acid or one of the metabolites produced is have self-catabolite repression [49]. Inhibition at higher pectin concentration in fermentative medium was reported in pectinase production by Aspergillus niger MTCC 281 by using natural substrates [50,51].
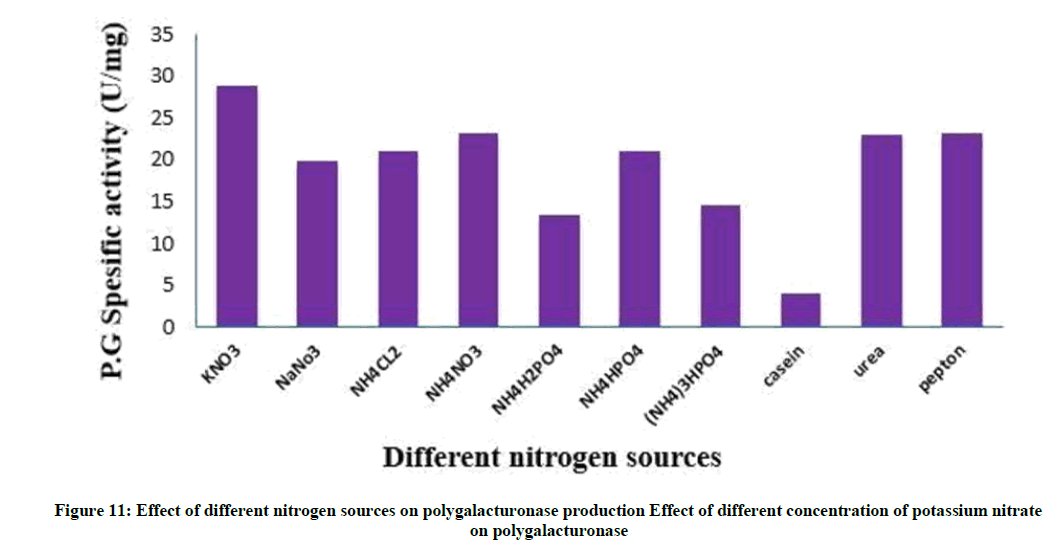
Nitrogen supplements, when included into the production medium, facilitate better biomass creation and subsequently higher metabolite secretion. Nitrogenous compounds are used by the microbial cells for the synthesis of nucleotides, amino acids, proteins, enzymes and other metabolites [36]. The effect of various organic and inorganic nitrogen supplements with the same equimolar N concentration of potassium nitrate was examined as shown in Figure 11.
Results reported that potassium nitrate supported maximum polygalacturonase synthesis. The low of enzyme productivity by other nitrogenous compounds may be resulted from the deviation of C/N ratio and subsequently higher carbon concentration obtained. Ammonium dihydrogen phosphate showed the lowest enzyme production, this may due to higher phosphorous concentration occur in fermentation medium hence lowered enzyme synthesis. Between Organic nitrogen supplements peptone showed good yield of polygalacturonase. This may be attributed to the nutritional vitamins and accessory growth factors naturally present in these organic supplements [36,37]. This results similar to [52] has potassium nitrate and peptone were the best nitrogen sources.
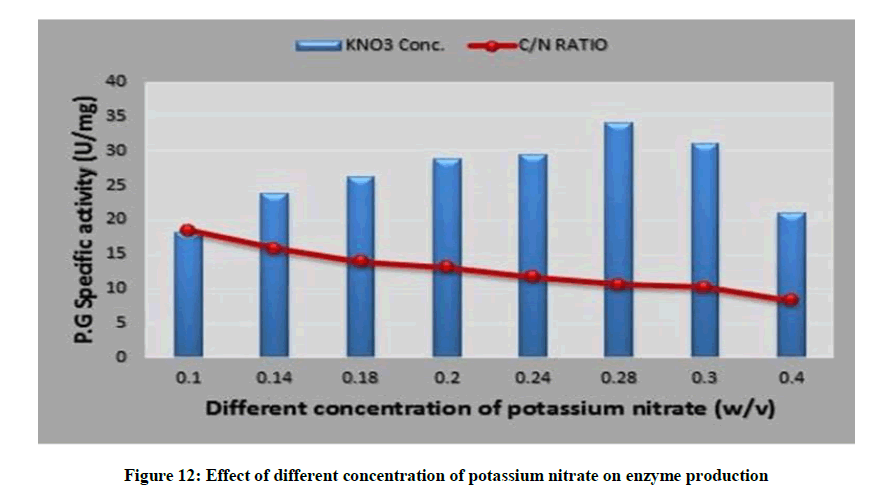
Interestingly, 0.28% potassium nitrate with (C/N=10.58) supported highest yield of polygalacturonase (34.09 U/mg), as depicted in Figure 12. Furthermore increase or decrease in the potassium nitrate percentage lead to deviation of (C/N) ratio rather than optimum one by thus leads to moderate level of the enzyme synthesis. A similar study of optimization of pectinase production by A. niger NCIM 548 reported maximal production (22.87 U/ml) at a C/N ratio of 5.9429 [53]. Also pectinase creation reported from Streptomyces sp. GHBA10 had highest pectinase production with a C/N ratio of 3.0 [37].
Conclusion
The current study focuses on molecular identification of Streptomyces sp. GHB5 and improvement of polygalacturonase production from this strain. Further studies well be done in the future in a large scale area.
References
- K. Mishra, S. Vuppu, S. Bishwambhar, Ind. J. Fund. Appl. Life Sci., 2011, 1, 4, 180-186.
- P. Sharma, N. Sharma, Int. J. Curr. Microbiol. App. Sci., 2018, 7, 1, 670-680.
- S.K. Anisa, S. Ashwini, K. Girish, Electronic J. Biol., 2013, 9, 2, 37-41.
- R. Daoud, M.K. Tahla, M.F. Azmeh, Int. J. Chem. Tech. Res., 2016, 9 1, 359-365.
- R.S. Jayani, S. Saxsena, R. Gupta, Proc. Biochem., 2005, 40, 2931-2944.
- P. Alimardani-Theuil, A. Gainvors-Claisse, F. Duchiron, Proc. Biochem., 2011, 46, 1525-1537.
- J.L. Adrio, A.L. Demain, Biomolecules., 2014, 4, 117-139.
- D. Fratebianchi, J.M. Crespo, C. Tarib, S. Cavalittoa, J. Chem. Tech. Biotech., 2016, 92, 305-310.
- N. Rajaei, N.S. Naghavi, K. Beheshtimaal, J. Sci., 2014, 11, 31, 109-115.
- F. Anwar, R. Naseer, M.I. Bhanger, S. Ashraf, F.N. Talpur, F.A. Aladedunye, J. Am. Oil Chem. Soc.,2008, 85, 321–330.
- Y.S. Huang, S.C. Ho, Food Chem., 2010, 119, 868–873.
- M.A. Martín, J.A. Siles, A.F. Chica, A. Martín, Bioresour. Technol., 2010, 101, 8993–8999.
- S.H. Handa, N. Sharma, S.h. Pathania, J. Ferment., 2016, 2, 2, 2-10.
- S. Palit, R. Banerjee, Biol. Technol., 2001, 44, 107–111.
- A.D. Juwon, O.F. Emmanuel, Biotech. Res. Int., 2012, 1-8.
- S.P. Lei, H.C. Lin, L. Heffernan, G. Wilcox, J. Bacteriol., 1985, 164, 2, 831-835.
- H.C.M. Kester, J. Visser, Biotechnol. & Appl. Biochem., 1990, 12, 150-160.
- C. Riou, G. Freyssinet, M. Feure, Appl. Environm. Microbiol., 1992, 58, 578-583.
- S.K. Sathish, P. Palanivelu, World J. of Microbiol & Biotech., 1998, 14, 781-782.
- E. Favela-Torres, T. Volke-Sepúlveda, G. Viniegra-Gonzalez, Food Tech. Biotech., 2006, 44, 221–227.
- S. Oncü, N. Gögus, C. Tari, S. Unluturk, F. Tokatli, Biochem. Eng. J., 2006, 32, 171–178.
- D.B. Pedrolli, A.C. Monteiro, E. Gomes, E.C. Carmona, The Open Biotech. J., 2009, 3, 9-18.
- M. Motalleb, H.A. Azad, M.R. Zamani, World Appl. Sci. J., 2008, 3, 96-101.
- T. Embley, E. Stackebrandt, Ann. Rev. Microbiol., 1994, 48, 257-289.
- Q. Y. M. Abdullah, M. A. Al-Maqtari, A A. Al-Awadhi, A. Y. Al-Mahdi, I. J. R. in Pharm. & Biosci.,2016,3, 3, 19-31.
- H. Lechevalier, M. Lechevalier, Ann. Rev. Microbiol., 1967, 21, 71-99.
- M. Goodfellow, S. Williams, Ecology of actinomycetes. Ann. Rev. Microbiol., 1983, 37, 186-216.
- J. Stach, A. Bull, Antonie van Leeuwenhoek., 2005, 87, 3-9.
- K. Wilkins, Chemosphere., 1996, 32, 1427-1434.
- S. Ballav, S.G. Dastager, S. Kerkar, Rec. Sci. Technol., 2012, 4, 31-39.
- N.H. Abd El-Nasser, E.S.M. El-Mahdy, W.G.H. Shosha, G.H. El Sayed, Res. J. Pharma., Biol. Chem. Sci.,2018, 9, 2, 744-751.
- K. Tamura, D. Peterson, N. Peterson, G. Stecher, M, Nei, S. Kumar, MEGA5: Mol. Biol. Evol.,2011, 28, 2731-2739.
- M. Bailey, P. Biely, K. Poutanen, J. Biotech., 1992, 23, 257–270.
- O.H. Lowry, N. J. Rosebrough, A. L. Farr, R. J. Randall, J. Biol. Chem., 1951, 193, 265-275.
- N. Kumar, Rec. res. In sci. & technol., 2010, 27, 36-42.
- J. Padhiar, A. Das, S. Bhattacharya, J. Biol. Sci., 2011, 14, 22, 1011-1018.
- D. Arijit, B. Sourav, R.V. Naimisha, S.S. Rajan, Inter. Res. J. Biol. Sci., 2013, 2, 3, 16-22.
- E. Venkata, N. Rajuand, G. Divakar, Internation. J. Pharma. Sci. & Res., 2016, 4, 2615-262.
- A. Kunamneni, K. Permaul, S. Singh, J. Biosci. Bioeng., 2005, 100, 168- 171.
- O. J. Oumer, D. Abate, Hindawi Enzy. Res., 2017, 1-7.
- S. Mrudula, R. Anitharaj, Global J. Biotech. & Biochem., 2011, 6, 64-71.
- R. Dhital, O.M.P. Panta, T.B. Karki, J. Food Sci. Tech. Nepal., 2013, 8, 65-70.
- R. Gupta and Kalpana, Acta Microbiol et Immunologica Hungarica., 2011, 58, 4, 339-349.
- P.B. Papdiwal, K.B. Deshpande, Arch. Biol. Chem., 2000, 7, 67-81.
- J.M. Willey, L.M. Sherwood, C.J. Woolverton, Prescott, Harley and Kleins Microbiology, 7th ed. McGraw Hill Co. Inc., Boston., 2008, 48.
- G.A. Aaisha, D.L. Barate, Int. J. Curr. Microbiol. App. Sci., 2016, 5, 1, 514-524.
- A. Kilara, Proc. Biochem., 1982, 17, 4, 35-41.
- E.V.N. Raju, G. Divakar, IJPSR, 2013, 4, 7, 2615-2622.
- S. Tsuymu, J. Bacteriol., 1979, 137, 1035-1036.
- M. Palaniyappan, V. Vijayagopal, R. Viswanathan, T. Viruthagiri, African j. Biotech., 2009, 8, 682-686.
- M. Joshi, M. Nerurkar, R. Adivarekar, Innovative Roman. Food Biotech., 2013, 12, 201.
- S.R. Jagiasi, National conference on Biodiversity, 'FAVEO' 2013, ISBN: 978-81-923628-1-6, 2013, 77-87.
- N.M. Yugandhar, D.V.R.R. Kumar, V. Prasanthi, N.K. Kumar, D.S.R. Reddy, Res. J. Microbiol.,2008, 3, 1, 9-16.