Research Article - Der Pharma Chemica ( 2018) Volume 10, Issue 8
Synthesis and Biological Evaluation of N,NÃÆâÃâââ¬Ãâââ¢-Bis(1-(2-Hydroxyphenyl)Ethylidene)-2-Methylmalonohydrazide SchiffÃÆâÃâââ¬Ãâââ¢s Base and its Metal Complexes of First Transition Series
Saini S1, Pal R1, Gupta AK1* and Beniwal V2
1Department of Chemistry, Maharishi Markandeshwar (Deemed To Be University), Mullana (Ambala), India
2Department of Biotechnology, Maharishi Markandeshwar (Deemed To Be University), Mullana (Ambala), India
- *Corresponding Author:
- Gupta AK
Department of Chemistry
Maharishi Markandeshwar (Deemed To Be University)
Mullana (Ambala), India
Abstract
In search of new chemical nucleases the synthesis of methylmalonohydrazide based hydrazone Schiff base and its colored complexes with transition metal Co(II), Ni(II), Cu(II), Mn(II), and Zn(II) have been synthesized. All synthesized compounds were screened for antibacterial activity against gram (+) and gram (-) bacteria using Oxacillin and Penicillin-G as a reference standard and their DNA photocleavage potential were carried out by using the plasmid DNA via gel electrophoresis method. From experimental results it has been observed that biological activity increases significantly upon coordination. The synthesis of hydrazone Schiff’s base, N,N’-bis(1-(2-Hydroxyphenyl)ethylidene)-2-methylmalonohydrazide (MAPLH2) have been achieved by a condensation reaction between 2-methylmalonohydrzide and 1-(2-Hydroxyphenyl)ethanone. Furthermore, new transition metal complexes were formed by the reaction between novel ligand and different transition metal ions in 1:1 stoichiometric ratio. All the synthesized compounds were characterized on the basis of various spectroscopic techniques like Proton Nuclear Magnetic Resonance (1H-NMR), Ultraviolet (UV)-Visible, elemental analysis, Fourier-transform infrared (FT-IR) and mass spectroscopy. 1H-NMR data of complexes showed the coordination of hydroxy atom with subsequent loss of proton and FT-IR spectral data reveals that the participation of nitrogen atom of ligand in coordination with metal ion.
Keywords
2-methylmalonohydrazide, 1-(2-Hydroxyphenyl)ethanone, Schiff’s base, Metal complexes, antibacterial activity, DNA photocleavage.
Introduction
There has been growing intensive interest in the topic of Schiff’s base metal complexes because of their various pharmaceutical [1] and biological [2,3] activities. Since they possess good chelating properties, it has been reported that the presence of -CO-NHN=C- moiety is mainly responsible for the biological activities of hydrazones [4]. It is well known that the presence of azomethine group (>C=N-) of Schiff’s bases made them important as a biologically active agents in medicinal chemistry [5]. The presence of >C=N- group also responsible for their antibacterial [6], antifungal [7], antituberculosis [8], antitumor [9], anticancer [10,11], antihelminthics [12], DNA cleavage [13] and DNA binding [14] properties. Complexation ability of many ligands with transition metal ions [15] were used for analytical applications such as quantitative determination of metal ion. It is also reported that Schiff’s bases were also used as catalyst in many reactions [16]. It is observed from the literature that the Schiff’s base based on 1-(2-Hydroxyphenyl)ethanone has been used to form a large number of heterocyclic derivatives that are known to possess potent antimicrobial and antifungal activity [17]. In view of above mentioned facts, we planned to synthesize a novel hydrazone Schiff’s base to synthesize some novel complexes. We synthesized Schiff’s base ligand by treatment of 2-methylmalonohydrazide with 1-(2-Hydroxyphenyl)ethanone. Finally synthesized ligand was treated with metal acetate salt of Cu(II), Ni(II), Co(II), Mn(II), and Zn(II) to form their respective Schiff’s base metal complexes. All the synthesized compounds were characterized on the basis of various technique like elemental analysis, Proton Nuclear Magnetic Resonance (1H-NMR), Ultraviolet (UV)-Visible spectroscopy, Fourier-transform infrared (FT-IR), mass spectroscopy etc. Moreover, Schiff’s base and its metal complexes were subjected to antibacterial and DNA photocleavage activity studies with an expectation to find some novel and potential antibacterial agents.
Materials and Methods
1-(2-Hydroxyphenyl)ethanone was purchased from Merck and used as such without purification. All other chemicals including solvents were of LR grade and were used as such without any purification or distillation. All the experiments were performed on magnetic stirrer fitted with hot plate under continuous stirring. Double distilled water was used in the present investigation. The ligand was purified from ethanol by recrystallization before preparing metal complexes. The 1H-NMR spectra of the ligand and its transition metal complexes were recorded on Bruker Advance II 400 MHz instrument at frequencies of 400 MHz and 100 MHz, respectively using TMS as an internal reference standard. IR spectra were recorded on Shimadzu IR Affinity in 4000-400 cm-1 range using KBr pellet method. The electronic spectra were recorded using Shimadzu UV 1800 instrument in DMSO as a solvent. Mass spectra were measured on Agilent Mass Spectrometer. Hydrogen, Carbon, Nitrogen and metal contents were analyzed using LECO 9320 analyzer. All the melting points were determined by using an open capillary method.
Synthesis of ligand
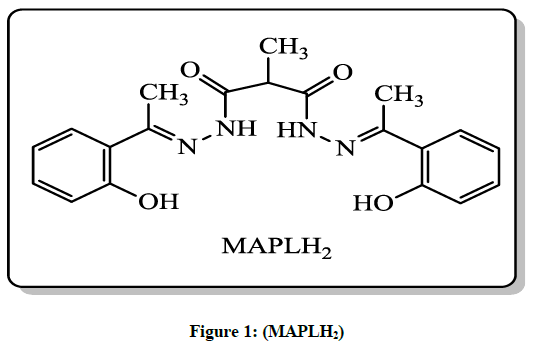
N,N’-bis(1-(2-Hydroxyphenyl)ethylidene)-2-methylmalonohydrazide (MAPLH2) (Figure 1)
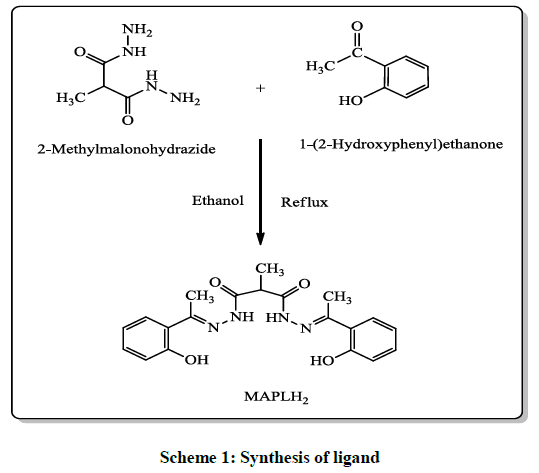
2-methylmalonohydrazide was prepared by reacting 2-methylmalonate (0.1 mol) and hydrazine hydrate (0.2 mol) in ethanol according to the literature procedure [18]. Ligand used in this research was prepared by the reaction of 2-methylmalonohydrazide (1.0 mol) with 1-(2- Hydroxyphenyl)ethanone (2.0 mol) according to the procedure mention below.
2-methylmalonohydrazide (1.0 mol) was dissolved in 40 ml ethanol (99.9%) in a 100 ml round-bottomed flask. The temperature of reaction mixture increased up to 80ºC, followed by dropwise addition of hot ethanolic solution of 1-(2-Hydroxyphenyl)ethanone (2.0 mol) to the above reaction mixture under continuous stirring. The reaction mixture maintained for 3.0 to 4.0 hours at 80ºC under continuous stirring. The progress of reaction was monitored by thin layer chromatography in methanol: chloroform system (5: 95). When reaction was completed, then reaction mass was cooled at room temperature. The off white crystal thus appeared were filtered out and washed 2-3 times with chilled EtOH. The crude product was recrystalized from EtOH and it was hold in vacuum dryer for 4-5 hours at 40°C-45°C to remove traces of solvent (Scheme 1). Product: Off white solid, Yield: 60%. M.p.:170ºC.
Preparation of N,N’-bis(1-(2-Hydroxyphenyl)ethylidene)-2-methylmalonohydrazide (MAPLH2)
Synthesis of metal complexes
General procedure
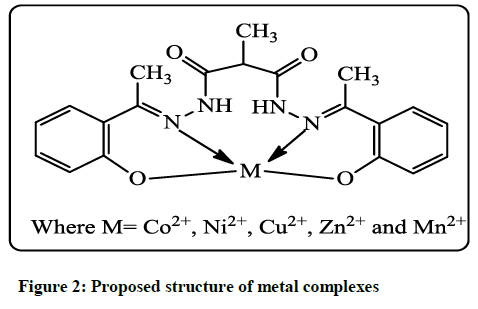
Various metal complexes were prepared by the treatment of Ligand MAPLH2 (0.1 mol) in 40 ml methanolic solution with appropriate metal salt (0.1 mol) dissolved in 15 ml of hot methanolic solution, in a 100 ml round bottom flask. It was heated with continuous stirring for 20 min to make a clear solution. The pH of reaction mixture was adjusted to eight by addition 10% NaOH in methanol solution. A clear solution thus obtained followed by reflux for 4 hours to prepare complexes with 1: 1 metals to ligand ratio. The precipitates of metal complexes thus formed were filtered out, without cooling and washed with methanol, finally washed twice with petroleum ether (nonpolar) solvent to get rid the traces of impurities. Complexes were dried in hot air oven at 60ºC-65ºC up to 4 hours. All transition metal complexes (Figure 2) were non hygroscopic and stable at room temperature.
(C20H22N4O4) Ligand
Off white in color, solid yield: 60%. M.p.: 170ºC; Molar conductance 7.5 Ohm-1 cm-1 mol-1; IR (KBr, cm-1) 3242(s), υ(O-H); 1568(s), υ(C=N); 1672(s), υ(C=O); 1232(s), υ(C-O).Electronic spectra (λmax nm) in DMSO: 264, 339. MS: m/z [M+1]+ 157.2, 382.4, 381.4. 1H NMR (DMSO-d6, 400 MHz) (ppm): 2.58-2.60 (s, 6H, -N=C-CH3); 2.19 (s, 3H, -CH3); 5.80 (s, 1H, -COCH3CHCO); 6.96-7.62 (s, 8H, Ar-H); 11.97 (s, 2H, -OH); 13.24 (s, 1H, -NH); Anal. Calculated for C20H22N4O4: C 62.82, H 5.80, N 14.65; found C 62.76, H 5.81, N 14.68.
(C20H20N4O4)Co
Light orange solid in color, yield: 62%. M.p.: 262ºC; Molar conductance 6.9 Ohm-1 cm-1 mol-1 IR (KBr, cm-1) 3238(s), υ(O-H); 1597(s), υ(C=N); 1672(s), υ(C=O); 1234(s), υ(C-O); 522(s), υ(M-N); 416(s), υ(M-O). Electronic spectra (λmax nm) in DMSO: 280, 327, 564. MS: m/z [M+1]+ 250.4, 259.2, 439.6, 446.6. Anal. Calculated for C20H20N4O4Co: C 54.68, H 4.49, N 12.75, M 13.41; found C 54.66, H 4.47, N 12.72, M 13.42.
(C20H20N4O4)Ni
Green in color, solid yield: 65%. M.p.: >290ºC; Molar conductance 8.0 Ohm-1 cm-1 mol-1; IR (KBr, cm-1) 3242(s), υ(O-H); 1566(s), υ(C=N); 1672(s), υ(C=O); 1232(s), υ(C-O); 522(s), υ(M-N); 414(s), υ(M-O). Electronic spectra (λmax nm) in DMSO: 285, 340, 557. MS: m/z [M+1]+ 231.2, 433.3, 439.5. Anal. Calculated for C20H20N4O4Ni: C 54.71, H 4.59, N 12.76, M 13.37; found C 54.73, H 4.59, N 12.75, M 13.35.
(C20H20N4O4)Cu
Green solid in color, yield: 70%. M.p.: >290ºC; Molar conductance 7.1 Ohm-1 cm-1 mol-1; IR (KBr, cm-1) 3242(s), υ(O-H); 1570(s), υ(C=N); 1672(s), υ(C=O); 1224(s), υ(C-O); 522(s), υ(M-N); 408(s), υ(M-O). Electronic spectra (λmax nm) in DMSO: 228, 270, 580. MS: m/z [M+1]+ 381.2, 382.2, 397.3, 443.2, 444.3, 445.9. Anal. Calculated for C20H20N4O4Cu: C 54.11, H 4.54, N 12.62, M 14.31; found C 54.10, H 4.54, N 12.65, M 14.33.
(C20H20N4O4)Mn
Yellow solid in color, yield: 60%. M.p.: 273ºC; Molar conductance 6.1 Ohm-1 cm-1 mol-1; IR (KBr, cm-1) 3240(s), υ(O-H); 1568(s), υ(C=N); 1672(s), υ(C=O); 1232(s), υ(C-O); 522(s), υ(M-N); 414(s), υ(M-O). Electronic spectra (λmax nm) in DMSO: 245, 306. MS: m/z [M+1]+ 247.5, 284.8, 436.6, 443.8, 457.8. Anal. Calculated for C20H20N4O4Mn: C 55.18, H 4.63, N 12.87, M 12.62; found C 55.10, H 4.60, N 12.88, M 12.65.
(C20H20N4O4)Zn
Colorless solid, yield: 70%. M.p.: 225ºC; Molar conductance 6.6 Ohm-1 cm-1 mol-1; IR (KBr, cm-1) 3192(s), υ(O-H); 1546(s), υ(C=N); 1660(s), υ(C=O); 1220(s), υ(C-O); 503(s), υ(M-N); 418(s), υ(M-O). Electronic spectra (λmax nm) in DMSO: 270, 335. MS: m/z [M+1]+ 157.2, 163.1, 167.1, 222.47, 229.9, 247.3, 263.3, 361.4, 363.1, 445.8. Anal. Calculated for C20H20N4O4Zn: C 53.89, H 4.52, N 12.57, M 14.67; found C 53.83, H 4.53, N 12.51, M 14.70.
Results and Discussion
Chemistry
In present study we report the synthesis of novel N,N’-bis(1-(2-Hydroxyphenyl)ethylidene)-2-methylmalonohydrazide (MAPLH2), is outlined in Scheme 1. Furthermore, its metal complexes were formed with first transition metal series. Ligand used in the present study was synthesized by condensation reaction between 2-methylmalonohydrazide and 1-(2-Hydroxyphenyl)ethanone. All the synthesized compounds were colored except Zn (II) complexes and non-hygroscopic in nature, stable at room temperature and were highly soluble in DMSO and DMF. The structures of synthesized compounds were established on basis of various analytical techniques like FT-IR, NMR, Mass spectrometry, UV, elemental analysis. The 1H-NMR chemical shift values of ligand (MAPLH2) and Zn complex are listed in Table 1. The 1H-NMR of free ligand and its metal complex with Zn were recorded by using dimethyl sulfoxide (DMSO-d6) solvent with tetramethylsilane (TMS) as an internal reference standard. The Synthesized compounds were further explored for in vitro DNA photocleaving as well as antibacterial activities.
| Proton | MAPLH2 1H (ppm) |
MAPLZn 1H (ppm) |
|---|---|---|
| -N=C-CH3 | 2.58-2.60, s, 6H | 1.88, s, 6H |
| CH (-COCH3CHCO) | 5.80, s, 1H | 3.56, s, 1H |
| CH3 (Malonohydrazine) | 2.19, s, 3H | 1.45-1.46, d, 3H |
| Ar-H | 6.96-7.62, 8H | 6.92-7.75, 8H |
| O-H (Phenolic) | 11.97, s, 2H | - |
| N-H | 13.24, s, 1H | 12.96, s, 1H |
Table 1: 1H-NMR of MAPLH2 and its MAPLZn complexes
FT-IR Spectroscopy
Infrared spectroscopy of free ligand is compared with metal complexes in order to explore the nature and mode of bonding of metal ions with ligand. FT-IR is a versatile technique to know the mode of binding as well as to find out the donor atoms of ligand involved in coordination with metal ions. The FT-IR spectrum of Schiff’s base ligand has been compared with that of metal complexes to know the binding mode. The FT-IR band observed in case of free ligand was at 3242 cm-1, 1568 cm-1, 1672 cm-1 and 1232 cm-1. This stretching frequencies may be observed due to υOH, υC=N, υC=O and υC-O respectively. All the above mentioned stretching frequencies were also assigned in case of metal complexes but with some change in frequencies which show involvement of phenolic oxygen atom in coordination with metal ions [19]. 1H-NMR data shows that the deprotonation of this phenolic oxygen atom takes place upon coordination with metal ions. The involvement of nitrogen atom of azomethine group in coordination with metal ions may easily be showed as υC=N stretch shift to different frequencies in the IR spectrum of metal complexes [20]. This change in frequencies may be due to donation of lone pair of electrons of nitrogen atom of azomethine to the empty d orbitals of metal ions. Additional stretching frequencies in case of metal complexes at 503-522 cm-1 and 408-418 cm-1 were assigned due to υM-N and υM-O which further confirmed the formation of metal complexes as well as participation of N, O atoms in coordination [21].
1H-NMR
The NMR spectroscopy is used to judge the exact chemical environment of organic molecules. The 1H-NMR of MAPLH2 and its metal complex with zinc(II) recorded in CDCl3 and deuterated dimethyl sulfoxide (DMSO-d6) respecyively, by using tetramethylsilane (TMS) as an internal reference standard are presented in Table 1. The proton NMR of MAPLH2 displayed signal at 2.19 ppm (doublet, 3H) due to -CH3 of malonohydrazine, 2.58-2.60 ppm (singlet, 6H) due to -CH3 of -N=C-CH3, 5.80 ppm (singlet, 1H) due to CO-CH3CH-CO, 6.96-7.62 ppm (8H) due to aromatic proton (Ar-H), 11.97 ppm (singlet, 2H) due to -OH of phenolic proton [22,23] and 13.24 ppm (singlet, 1H) due to -NH proton. The proton NMR spectra of MAPLZn complex showed the deprotonation of phenolic proton because disappearances of phenolic proton signal which further indicate the coordination of metal ions through phenolic oxygen of hydrazone. Other recorded chemical shift values displayed the signal at 1.45-4.46 ppm (doublet, 3H) due to -CH3 of malonohydrazine, 1.88 ppm (singlet, 6H) due to -CH3 of -N=C-CH3, 3.56 ppm (singlet, 1H) due to CO-CH3CH-CO, 6.92-7.75 ppm (8H) due to aromatic proton, 12.96 ppm (singlet, 1H) due to -NH proton as shown in Table 1.
UV-VIS spectra
The UV-visible spectra of the MAPLH2 and all synthesized metal complexes were analyzed in 10-4 M dimethyl sulfoxide solution at RT as shown in Table 2. The UV-VIS spectrum of MAPLH2 displays two signals at 264 and 339 nm. These absorption band were appeared, because the presence of π-π* and n-π* transition of –C=N group and -C=O moiety [24]. These UV absorption bands were also present in case of synthesized metal complexes but with hypsochromic shift in comparison to MAPLH2, suggesting the participation of -C=N group in coordination with metal ions. The MAPLCo, MAPLNi and MAPLCu showed band center at 564, 557 and 580 nm, respectively due to d-d transition. This proposed the square planner geometry over the respective metal ions.
| Compounds | IR frequency (Stretching’s in cm-1) | λmax (nm) | |||||
|---|---|---|---|---|---|---|---|
| υO-H | υC=N | υC=O | υC-O | υM-N | υM-O | ||
| MAPLH2 | 3242 | 1568 | 1672 | 1232 | - | - | 264, 339, |
| MAPLCo | 3238 | 1597 | 1672 | 1234 | 522 | 416 | 280, 327, 564 |
| MAPLNi | 3242 | 1566 | 1672 | 1232 | 522 | 414 | 285, 340, 557 |
| MAPLCu | 3242 | 1570 | 1672 | 1224 | 522 | 408 | 270, 228, 580, |
| MAPLMn | 3240 | 1568 | 1672 | 1232 | 522 | 414 | 245, 306 |
| MAPLZn | 3192 | 1546 | 1660 | 1220 | 503 | 418 | 270, 335 |
Table 2: IR and UV data of ligand and its metal complexes
Mass spectroscopy
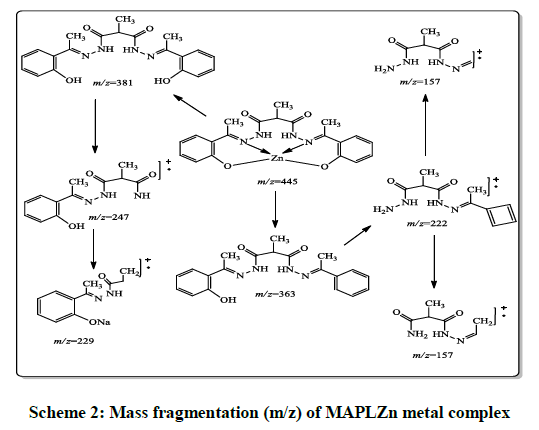
The mass fragmentation data of Schiff’s base ligand as well as its transition metal complexes are presented in Tables 3 and 4. Mass of ligand and its metal complexes were confirmed on the basis of mass spectroscopy. The mass spectra of hydrazone (MAPLH2) display an intense molecular ion peaks at m/z 382.4, 381.4 and 157.2 suggested the formation of hydrazone. The mass spectrum of MAPLCo, MAPLNi, MAPLCu, MAPLMn and MAPLZn display molecular ion peaks at m/z 439.6, 439.5, 444.3, 436.6 and 445.9, respectively. The mass fragmentation (m/z) (Scheme 2) results give strong clue about the formation of metal complex in 1: 1 metal to hydrazone ratio. The metal complexes results showed all the mass fragments observed in case of free ligand.
| Compounds | Fragmentation Value (m/z) |
|---|---|
| MAPLH2 | 382.4, 381.4 and 157.2. |
| MAPLCo | 439.6, 446.6, 259.2 and 250.4. |
| MAPLNi | 439.5, 433.3 and 231.2. |
| MAPLCu | 445.9, 444.3, 443.2, 397.3, 382.2 and 381.2. |
| MAPLMn | 443.48, 457.8, 436.6, 284.8 and 247.5. |
| MAPLZn | 445.8, 363.1, 361.4, 263.3, 247.3, 229.9, 222.47, 167.1, 163.1 and 157.2 |
Table 3: Mass fragmentation (m/z) data of ligand and its metal complexes
| Compounds | Colour | MP (ºC) | Found (Calculated)% | Conductance Ohm-1 cm2 mol-1 | |||
|---|---|---|---|---|---|---|---|
| C | H | N | M | ||||
| MAPLH2 | Off White | 170 | 62.76 | 5.81 | 14.68 | - | 7.5 |
| -62.82 | -5.8 | -14.65 | |||||
| MAPLCo | Orange | 262 | 54.66 | 4.47 | 12.72 | 13.42 | 6.9 |
| -54.68 | -4.49 | -12.75 | -13.41 | ||||
| MAPLNi | Green | >290 | 54.73 | 4.59 | 12.75 | 13.35 | 8 |
| -54.71 | -4.59 | -12.76 | -13.37 | ||||
| MAPLCu | Green | >290 | 54.1 | 4.54 | 12.65 | 14.33 | 7.1 |
| -54.11 | -4.54 | -12.62 | -14.31 | ||||
| MAPLMn | Yellow | 273 (d) | 55.1 | 4.6 | 12.88 | 12.65 | 6.1 |
| -55.18 | -4.63 | -12.87 | -12.62 | ||||
| MAPLZn | Colorless | 225 | 53.83 | 4.53 | 12.51 | 14.7 | 6.6 |
| -53.89 | -4.52 | -12.57 | -14.67 | ||||
Table 4: Physio-chemical data of Schiff’s base MAPLH2 and its metal complexes
Biological activity
DNA photocleavage activities
DNA photocleavage cleavage reaction of transition metal complexes have been investigated in recent years. DNA is a target molecule for most of anticancer drugs, in past decade extensive study has been done to explore the results of interactions between small molecules like coordinated metal complexes with DNA molecule. This area of research is much more important as it can provide some effective and efficient solution against life threatening diseases like cancer. In our present study, DNA photocleavage study on plasmid DNA was performed by gel electrophoresis technique. The result of DNA photocleavage study is showed in Figure 3. DNA photocleavage reaction was performed between synthesized compounds and DNA. The prepared solution was diluted with loading dye using 1% agarose gel. Ethidium bromide was added to the above prepared solution and mixed well. The solution of agarose was poured on glass tray and clamped immediately with comb to form sample wells. The gel was placed in electrophoresis tank, sufficient amount of electrophoretic buffers were added to cover the gel. The plasmid DNA sample, metal complexes and buffer were mixed with loading dye and loaded into the well of gel by using micropipette. The electrical current pass through running buffer, after 2-3 h, the gel was taken out of buffer. After complete electrophoresis reaction the gel was photographed under UV transilluminator. From Figure 3 it is clear that control (C) consist of supercoiled (SC) and open circular forms (OC) of plasmid DNA, performed by gel electrophoresis technique. In case of MAPLCu (lane 3rd) there is fully transformation of SC form into nicked coiled (NC) and open circular (OC) [25] forms which revealed that the breakage of one strand of SC form of DNA. Whereas, MAPLH2 (lane 1st), MAPLCo (lane 2nd), MAPLNi (lane 4th), MAPLMn (lane 5th) and MAPLZn (lane 6th) showed that there is formation of supercoiled (SC) as well as open circular form of plasmid DNA. Furthermore, in all synthesized compounds there was a formation of smear from lane 1st to lane 7th. This observation suggested that Cu(II) possessed a good level of DNA photocleavage activity under similar experimental condition. However, all other compounds possess moderate DNA photo cleaving activity. Moreover, DNA cleaving capacity of metal complexes is increases in comparison to hydrazone (MAPLH2) because non covalent interaction of synthesized complexes with plasmid DNA consequently binding with DNA and finally DNA cleavage. The difference in the cleavage potential of compounds was due to different extent of binding ability of different metal complexes.
Lane C (DNA+dimethyl sulfoxide), lane 1st (DNA+MAPLH2), lane 2nd (DNA+MAPLCo), lane 3rd (DNA+MAPLCu), lane 4th (DNA+MAPLNi), lane 5th (DNA+MAPLMn) and lane 6th (DNA+MAPLZn).
Antibacterial activity of MAPLH2 and its complexes
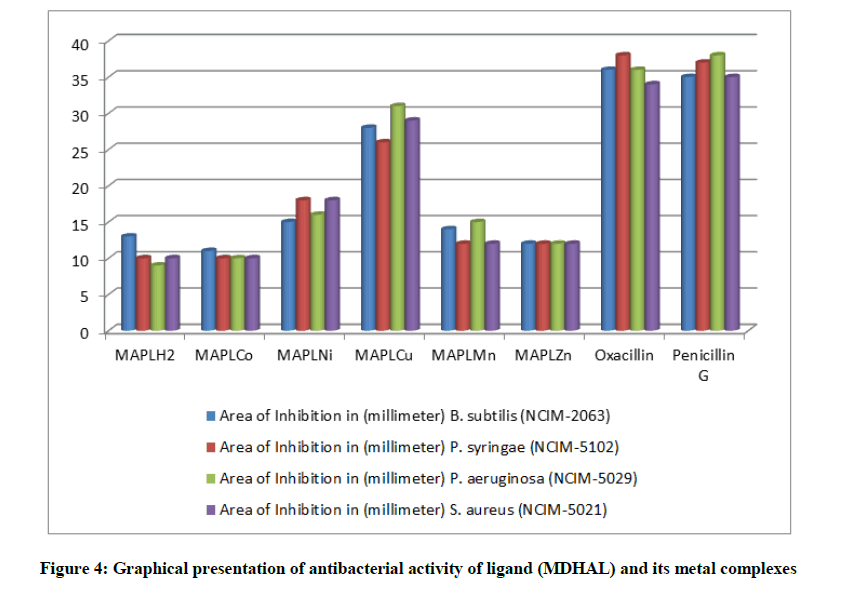
The in vitro antibacterial activity of synthesized Schiff’s base ligand and its metal complexes of first transition series have been tested against two gram positive bacteria B. subtillis, S. aureus, and two gram negative bacteria P. syringae, P. aeruginosa by well diffusion method. The antibacterial activity of these compounds was compared with Oxacillin and Penicilline G as a standard reference. The antibacterial activity obtained for the synthesized compounds are listed in Figure 4 and Table 5. All the synthesized compounds showed a significantly antibacterial activity against bacteria. It has been found that the activity is remarkably increased on coordination. Furthermore, polarity of the metal ion reduces on coordination [26] and this cause increase in the lipophilic character of metal chelate so as to make it more permeable through the lipid layer of microorganism [27] thus destroying them more effectively. The anti-bacterial activity of MAPLH2 and its complexes is found to be in the order MAPLCu>MAPLNi>MAPLMn>MAPLZn>MAPLCo>MAPLH2. The obtained result reflect the higher activity of Cu(II) complexes can be explained as, on chelation the polarity of Cu(II) reduced to a greater extent due to effective partial sharing of the positive charge of the copper ion by the donor atoms of ligand groups. Therefore, Cu(II) ions can easily penetrate and inhibit their growth more effectively the cell wall of microorganisms [28,29] as compared to ligand (MAPLH2). On chelation with metal ions the delocalization of p-electrons over the fully chelate ring will be increased which increases the penetration power of the complex into lipid membranes and blocking the metal binding sites in the enzyme of microorganism [30].
| Compounds | Area of Inhibition in (mm) | |||
|---|---|---|---|---|
| Bacillus subtilis (NCIM-2063) |
Pseudomonas syringae (NCIM-5102) |
Pseudomonas aeruginosa (NCIM-5029) |
Staphylococcus aureus (NCIM-5021) |
|
| MAPLH2 | 13 | 10 | 9 | 10 |
| MAPLCo | 11 | 10 | 10 | 10 |
| MAPLNi | 15 | 18 | 16 | 18 |
| MAPLCu | 28 | 26 | 31 | 29 |
| MAPLMn | 14 | 12 | 15 | 12 |
| MAPLZn | 12 | 12 | 12 | 12 |
| Oxacillin | 36 | 38 | 36 | 34 |
| Penicillin G | 35 | 37 | 38 | 35 |
Table 5: Anti-bacterial activity data of MAPLH2, MAPLCo, MAPLNi, MAPLCu, MAPLMn and MAPLZn
Conclusion
In our research, we report the syntheses, characterization and biological activity of MAPLH2 and its complexes by treating it with metal acetate of first transition series. Spectral data revealed the involvement of -N atom of -C=N and -O atom of hydroxyl group in coordination with metal ions. Metal to ligand stoichiometric ratio (1:1) has been proved on the basis of mass spectrometry and elemental analysis. It was observed that MAPLCo, MAPLNi, MAPLCu, MAPLMn and MAPLZn possess excellent DNA photo cleaving activity in comparison to MAPLH2. In vitro antibacterial studies showed that activity enhanced remarkably on coordination with metal ions in comparison to free ligand under similar experimental condition. It was observed that, MDHALCu and MDHALNi complexes possess good level of anti-bacterial activity against the tested strains of bacteria. Structural modification may leads to same effective chemical nucleases as well as antibacterial agent in future.
Acknowledgement
The authors would like to acknowledge Maharishi Markandeshwar (Deemed to be University), Mullana (Ambala), India for providing the requisite facilities for the said work.
References
- O.D. Can, M.D. Altintop, U.D. Ozkay, U.I. Ucel, B. Dogruer, Arch. Pharm. Res., 2012, 35, 659.
- S. Saini, A.K. Gupta, R.P. Saini, R. Kumar, V. Beniwal, W.J.P.P.S., 2014,3, 1621-1636.
- D.P. Singh, V. Malik, R. Kumar, K. Kumar, J. Serb. Chem. Soc., 2010, 75, 763-772.
- A.A. Al-Amiery, Y.K. Al-Majedy, H.H. Ibrahim, A.A. Al-Tamimi, Org. Med. Chem. Lett., 2012, 2, 4.
- S.N. Pandeya, D. Sriram, G. Nath, E. De Clercq, Eur. J. Pharma. Sci., 1999, 9, 25-31.
- K. Anuradha, R. Rajavel, Int. J. Pharm. Technol., 2011, 3, 2217-2227.
- D.P. Singh, V. Malik, R. Kumar, K. Kumar, J. Serb. Chem. Soc., 2010, 75, 763-772.
- S.A. Patila, M. Manjunatha, U.V. Kamblea, P.S. Badami, Der. Pharma. Chemica, 2011, 3, 97-108.
- E.W. Ainscough, A.M. Bordie, A.J. Dobbs, J.D. Ranford, J.M. Waters, Inorg. Chim. Acta., 1998, 267, 27-38.
- A.A. Osowole, I. Ott, O.M. Ogunlana, Int. J. Inorg. Chem., 2012, 1-6.
- M.I. Husain, M.A. Shukla, S.K. Agarwal, J. Ind. Chem. Soc., 1979, 56, 306-307.
- R.P. Saini, V. Kumar, A.K. Gupta, G.K. Gupta, Med. Chem. Res., doi: 10.1007/s00044-013-0657-6, 2013.
- A.D. Kulkarni, S.A. Patil, V.H. Naik, P.S. Badami, Med. Chem. Res., 2011, 20, 346-354.
- M.R. Ganjali, P. Norouzi, T. Alizadeh, N.M. Salavati, Bull. Korean Chem. Soc., 2007, 28, 1.
- K.C. Gupta, A.K. Sutar, Coord. Chem. Rev., 2008, 252, 1420-1450.
- C. Redshaw, Catalysts, 2017, 7, 1-11.
- M. Tyagi, S. Chandra, P. Tyagi, J. Akhtarb, A. Kandanb, B. Singh, J. Taibah. Univ. Sci., doi: 10.1016/j.jtusci.2015.11.003, 2016.
- R. Rajavel, M.S. Vadivu, C. Anitha, ECHEM., 2008, 5(3), 620-626.
- S. AbouEl-Enein, F.A. El-Saied, S.M. Emam, M.A. Ell- Salamony M.A., Spec. Chim. Acta, 2008, 71, 421-429.
- M.A. Phaniband, S.D. Dhumwad, S.R. Pattan, Med. Chem. Res., 2011, 20, 493-502.
- K. Nakamoto, Wiley, New York, 1986.
- N.P. Singh, V.P. Tyagi, B. Ratnam, J. Chem. Pharm. Res., 2010, 2, 473-477.
- Y.T. Liu, G.D. Lian, D.W. Yin, B.J. Su, Spectrochim. Acta, Part A., 2013, 100, 131-137.
- R.S. Joseyphus, C.J. Dhanaraj, M.S. Nair, Transition Met. Chem., 2006, 31, 699-702.
- Q.L. Zhang, J.G. Liu, J.Z. Liu, H. Li, Y. Yang, H. Xu, H. Chao, L.N. Ji, Inorg. Chim. Acta., 2002, 34, 339.
- A.B.P. Lever, Inorg. Electronic. Spectrosc., Elsevier, 1984.
- U.M. Hassan, Z.H. Chohan, C.T. Supuran, Main Group Met. Chem., 2002, 25, 291-296.
- M.B. Ferrari, S. Capacchi, G. Reffo, P. Tarasconi, R. Albertini, S. Pinelli, P. Lunghi, Inorg. Chim. Acta, 1999, 286, 134-141.
- Z.H. El-Wahab, M.M. Mashaly, A.A. Salman, B.A. El-Shetary, A.A. Faheim, Spec. Chim. Acta, 2004, 60, 2861-2873.
- M.A. Abdel-Nasser, Alaghaz, E.Z. Mohamed, A. Suliman, Alharbi, A.A. Reda, Ammar, J. Mol. Struct., 2015, 1087, 60-67.