Research Article - Der Pharma Chemica ( 2018) Volume 10, Issue 8
One Pot Multicomponent Synthesis, Biological and DNA Interactions of Cyclohepta[B]Indole Derivatives
Prakash B1, Amuthavalli A1, Edison D2, Sivaramkumar MS1, Kirubavathy SJ3 and Velmurugan R1*
1Department of Chemistry, Kongunadu Arts and Science College, Coimbatore-641029, Tamil Nadu, India
2Department of Science and Humanities, Adithya Institute of Technology, Coimbatore-641107 Tamil Nadu, India
3Department of Chemistry, PSGR Krishnammal College for Women, Coimbatore-641004, Tamil Nadu, India
- *Corresponding Author:
- Velmurugan R
Department of Chemistry
Kongunadu Arts and Science College
Coimbatore-641029, Tamil Nadu, India
Abstract
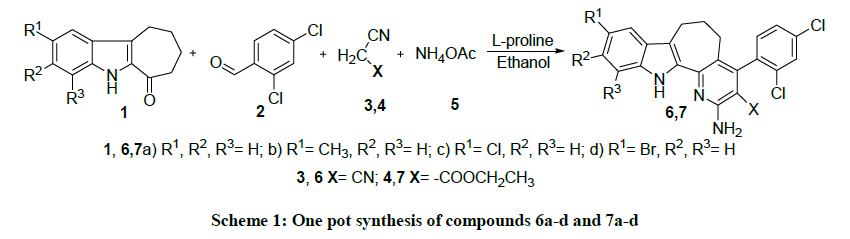
A chronological one-pot, atom economical multicomponent reaction yielding biologically capable cyclohepta[b]indole derivatives (6a-d, 7a-d) through a tandem Knoevenagel condensation followed by Michael addition. The reaction takes place under the organocatalyst L-proline. All the compounds were characterized by spectral and elemental analysis. The compounds were tested for their antimicrobial and DNA binding/cleavage studies. Antimicrobial results indicated that all the compounds have been significantly inhibit the growth of tested microorganisms. Especially compounds with chloro group exhibit potent activity. DNA cleavage of the compounds was study by agarose gel electrophoresis method with pUC19 DNA and found that most of the compounds have significant ability to cleave the DNA. Furthermore, the DNA binding property of the potent derivatives (6c, 7c) was studied by absorption spectra, the results showed that these compounds binds to calf thymus DNA (CT-DNA) in an intercalative mode and its intrinsic binding constants Kb is 3.74 × 104 and 6.73 × 104.
Keywords
Multicomponent, Cyclohepta[b]indole, Antimicrobial, DNA binding/cleavage.
Introduction
Multicomponent one pot reactions have huge attention in the synthesis of biologically efficient organic compounds using readily available starting materials with step ecomomics condition is the everlasting aspire of synthetic organic and medicinal chemistry [1,2]. This is due to its capability to carrying simultaneously more than three components in one pot, yielding highly proficient molecules and high atom economy [3-5]. Many of the biologically active different heterocycles systems are prepared by using one pot multicomponent reactions [6-9].
Nitrogen heterocycles in general and indoles in particular, are important scaffolding in biology. Indole and its derivatives are most attracting remarkable consideration because of their vast range of pharmaceutical and medicinal activities, numerous occurrences in natural products [10,11]. A huge number of synthetic indoles have been found an admired significance in pharmaceutical and medical applications, since they can able to bind to many receptors with high affinity [12-14]. Among them cyclohepta[b]indole derivatives have great attention because they exist in many natural products such as methuenine, ajmaline, 20-epiervatamine, ervitsine, episilicine, homoarcyriaflavin and caulersine [15,16] and it often shows broad range of potent biological activities, for example anticancer, antimicrobial, antitumor, anti-inflammatory and antimalarial activities [17,18].
Inspired by the biological significance of cyclohepta[b]indole derivatives and in continuation of our work, we employed multicomponent one pot synthesis of biologically active cyclohepta[b]indole derivative via Knoevenagel condensation reaction using a organocatalyst L-proline and the derivatives were characterized by Fourier-Transform Infrared Spectroscopy (FT-IR), Proton Nuclear Magnetic Resonance (1H-NMR), Carbon-13 Nuclear Magnetic Resonance (13C-NMR) and elemental analysis. The pharmacological activities of the synthesized compounds were evaluated by their antimicrobial, DNA cleavage and DNA binding properties.
Materials and Methods
Experimental section
General
Melting points (degree celsius) of the synthesized compounds were checked in open capillary tubes by using a Raaga melting point apparatus and found uncorrected. All the chemicals and solvents were purchased from Sigma-Aldrich and Merck, India. All reactions were carried out under atmospheric air and the products were checked by thin layer chromatography on TLC silica gel 60 F254 using eluting solvents such as ethyl acetate and petroleum ether. The synthesized compounds were purified by column chromatography using column silica gel 100-200 mesh (Petroleum ether /ethyl acetate 1: 2). FT-IR (KBr, cm-1) spectra were recorded on a Shimadzu spectrophotometer in the region of 4000–400 cm−1. 1H-NMR spectra were recorded on Bruker Avance III Spectrometer (400 MHz) with the solvent DMSO. 13C-NMR spectra were recorded on Brucker Avance 300 Spectrometer in CDCl3 (75 MHz) using tetramethylsilane (TMS) as internal standard. Coupling constants (J values) are reported in Hertz. Elemental analyses were performed on an Elementar Vario EL III CHN analyser.
Chemical synthesis
Synthesis of cyclohepta[b]indoles (6a-d)
A mixture of 7,8,9,10-Tetrahydro-5H-cyclohepta[b]indol-6-one derivatives (1a-d, 1 mmol), 2,4-dichlorobenzaldehyde (2, 1 mmol), malononitrile (3, 1 mmol), ammonium acetate (5, 1 mmol) and L-proline (1 mmol) in ethanol (20 ml) was refluxed on a oil bath for 2 h. After completion of the reaction, the excess of solvent was removed. The precipitate formed was dissolved in ice water and extracted with ethyl acetate. Combined organic layers were dried over anhydrous sodium sulfate. It was then purified by column chromatography over silica gel using petroleum ether: ethyl acetate (99: 1) as eluent to yield the respective compounds 6a-d.
2-Amino-4-(2,4-dichloro-phenyl)-5,6,7,12-tetrahydro-1,12-diaza-dibenzo[a,e]azulene-3-carbonitrile (6a): Yield; 65%; M.p. 223ºC-225ºC; FT-IR (cm-1): 3401 (N-H), 3325, 3251(NH2), 2207 (C≡N), 1612 (C=N); 1H-NMR (400 MHz, DMSO) δ (ppm)=2.37 (p, 2H, J= 2.4 Hz); 2.88 (t, 2H, J=5.2 Hz); 3.27 (t, 2H, J=6.4 Hz); 4.42 (s, 2H, NH2); 7.08-7.56 (m, 7H, Ar-H); 11.45 (s, 1H, indole-NH); 13C-NMR (100 MHz, CDCl3) δ (ppm)=26.72, 27.53, 31.49, 91.84, 114.49, 116.09, 120.19, 120.53, 121.12, 123.35, 123.41, 123.58, 124.85, 126.14, 127.21, 133.52, 133.89, 134.43, 136.14, 137.21, 153.52, 165.19, 167.43. Anal. Calcd For C23H16N4Cl2 (419.30): C, 65.88, H, 3.84, N, 13.36; Found: C, 65.89, H, 3.86, N, 13.38.
2-Amino-4-(2,4-dichloro-phenyl)-9-methyl-5,6,7,12-tetrahydro-1,12-diaza-dibenzo[a,e]azulene-3-carbonitrile (6b): Yield; 68%; M.p. 227ºC-229ºC; FT-IR (cm-1): 3417 (N-H), 3347, 3275 (NH2), 2218 (C≡N), 1657 (C=N); 1H-NMR (400 MHz, DMSO) δ (ppm)=2.12 (p, 2H, J=5.6 Hz); 2.39 (s, 3H, CH3); 2.82 (t, 2H, J=5.2 Hz); 3.11 (t, 2H, J=6.4 Hz); 4.43 (s, 2H, NH2); 7.13-7.65 (m, 6H, Ar-H); 11.26 (s, 1H, indole-NH); 13C-NMR (100 MHz, CDCl3) δ (ppm)=11.50, 15.47, 22.46, 27.08, 90.47, 114.52, 117.50, 120.01, 121.45, 123.62, 124.96, 125.41, 125.57, 126.84, 127.14, 127.96, 129.22, 129.67, 134.31, 135.11, 137.72, 151.88, 164.21, 168.14. Anal. Calcd. For. C24H18N4Cl2 (433.33): C, 66.52, H, 4.18, N, 12.92; Found: C, 66.55, H, 4.16, N, 12.94.
2-Amino-9-chloro-4-(2,4-dichloro-phenyl)-5,6,7,12-tetrahydro-1,12-diaza-dibenzo[a,e]azulene-3-carbonitrile (6c): Yield; 85%; M.p. 288ºC -291ºC; FT-IR (cm-1): 3392 (N-H), 3316, 3243 (NH2), 2245 (C≡N), 1645 (C=N); 1H-NMR (400 MHz, DMSO) δ (ppm)=2.73 (t, 2H, J=7.2 Hz); 2.90 (t, 2H, J=4 Hz); 3.79 (p, 2H, J=6.8 Hz); 4.01 (s, 2H, NH2); 7.19-7.24 (m, 6H, Ar-H); 10.89 (s,1H, indole-NH); 13C-NMR (100 MHz, CDCl3) δ (ppm)=26.84, 31.45, 34.79, 94.49, 115.54, 117.34, 122.12, 123.34, 123.75, 124.42, 124.83, 125.24, 126.42, 128.84, 132.98, 134.23, 134.96, 135.94, 136.16, 136.64, 155.46, 161.93, 169.52. Anal. Calcd. For C23H15N4Cl3 (453.75): C, 60.88, H, 3.33, N, 12.34; Found: C, 60.89, H, 3.36, N, 12.36.
2-Amino-9-bromo-4-(2,4-dichloro-phenyl)-5,6,7,12-tetrahydro-1,12-diaza-dibenzo[a,e]azulene-3-carbonitrile (6d): Yield; 78%; M.p. 264ºC -266 ºC; FT-IR (cm-1): 3423 (N-H), 3371, 3291 (NH2), 2231 (C≡N), 1623 (C=N); 1H-NMR (400 MHz, DMSO) δ (ppm)=2.17 (p, 2H, J=2.4 Hz); 2.88 (t, 2H, J=5.6 Hz); 3.15 (t, 2H, J=6.4 Hz); 4.05 (s, 2H, NH2); 7.09-7.80 (m, 6H, Ar-H); 11.64 (s,1H, indole-NH); 13C-NMR (100 MHz, CDCl3) δ (ppm)=21.56, 22.96, 24.30, 88.24, 113.27, 117.56, 120.24, 120.35, 122.63, 123.88, 125.22, 126.74, 126.92, 128.84, 132.22, 134.46, 135.23, 135.94, 137.65, 139.43, 156.25, 163.14, 164.19. Anal. Calcd For C23H15N4Cl2Br (498.20): C, 55.44, H, 3.03, N, 11.24; Found: C, 55.46, H, 3.06, N, 11.26.
General procedure for the ynthesis of cyclohepta[b]indoles (7a-d)
A mixture of 7,8,9,10-Tetrahydro-5H-cyclohepta[b]indol-6-one derivatives (1a-d, 1 mmol), 2,4-dichlorobenzaldehyde (2, 1 mmol), ethyl cyanoacetate (4, 1 mmol), ammonium acetate (5, 1 mmol) and L-proline (1 mmol) in ethanol (20 ml) was refluxed on a oil bath for 2 hr. The reaction was monitored by TLC, after the completion of the reaction the excess solvent was removed. The precipitate formed was dissolved in ice water and extracted with ethyl acetate. Combined organic layers were dried over anhydrous sodium sulfate. It was then purified by column chromatography over silica gel using petroleum ether: ethyl acetate (98:2) as eluent to yield the respective compounds 7a-d.
2-Amino-4-(2,4-dichloro-phenyl)-5,6,7,12-tetrahydro-1,12-diaza-dibenzo[a,e]azulene-3-carboxylic acid ethyl ester (7a): Yield; 72%; M.p. 256ºC -257 ºC; FT-IR (cm-1): 3425 (N-H), 3401, 3341 (NH2), 1695 (C=O), 1635 (C=N); 1H-NMR (400 MHz, DMSO) δ (ppm)= 1.73 (t, 3H, J=7.2 Hz); 2.16 (p,2H, J= 3.2 Hz); 3.13 (t, 2H, J=6.4 Hz); 3.26 (t, 2H, J=2.8 Hz); 4.67 (s, 2H, NH2); 5.05 (q, 2H, OCH2CH3, J=7.6 Hz); 7.28-7.70 (m, 7H, Ar-H); 11.48 (s,1H, indole-NH); 13C-NMR (100 MHz, CDCl3) δ (ppm)=15.49, 17.57, 19.76, 25.85, 49.26, 103.93, 112.70, 113.41, 122.26, 123.49, 125.15, 126.77, 127.45, 128.58, 130.32, 132.63, 134.76, 136.62, 137.45, 137.88, 138.32, 156.31, 158.76, 168.66, 169.32. Anal. Calcd For C25H21N3O2Cl2 (466.35): C, 64.38, H, 4.53, N, 9.01; Found: C, 64.39, H, 4.56, N, 9.03.
2-Amino-4-(2,4-dichloro-phenyl)-9-methyl-5,6,7,12-tetrahydro-1,12-diaza-dibenzo[a,e]azulene-3-carboxylic acid ethyl ester (7b): Yield; 66%; M.p. 248ºC -250 ºC; FT-IR (cm-1): 3428 (N-H), 3407, 3375 (NH2), 1710 (C=O), 1632 (C=N); 1H-NMR (400 MHz, DMSO) δ (ppm)= 1.30 (t, 3H, J=8 Hz); 1.85 (t, 2H, J=6 Hz); 2.05 (p,2H, J= 10.4 Hz); 2.47 (s, 3H, CH3); 2.95 (t, 2H, J=10 Hz); 3.99 (s, 2H, NH2); 4.31 (q, 2H, OCH2CH3, J=6.4 Hz ); 7.20-7.83 (m, 6H, Ar-H); 11.80 (s,1H, indole-NH); 13C-NMR (100 MHz, CDCl3) δ (ppm)= 15.56, 17.26, 18.82, 22.71, 27.32, 54.06, 104.07, 113.32, 114.48, 123.59, 124.53, 125.80, 126.19, 126.92, 127.50, 128.38, 129.47, 130.28, 130.90, 136.56, 137.35, 138.38, 154.47, 156.68, 160.53, 167.36. Anal. Calcd For C26H23N3O2Cl2 (480.38): C, 65, H, 4.82, N, 8.74; Found: C, 65.02, H, 4.83, N, 8.73.
2-Amino-9-chloro-4-(2,4-dichloro-phenyl)-5,6,7,12-tetrahydro-1,12-diaza-dibenzo[a,e]azulene-3-carboxylic acid ethyl ester (7c): Yield; 76%; M.p. 269ºC -271 ºC; FT-IR (cm-1): 3437 (N-H), 3398, 3257 (NH2), 1715 (C=O), 1641 (C=N); 1H-NMR (400 MHz, DMSO) δ (ppm)=1.79 (t, 3H, J=6.8 Hz); 2.20 (p,2H, J= 8 Hz); 2.35 (t, 2H, J=2.4 Hz); 2.85 (t, 2H, J=6 Hz); 3.87 (s, 2H, NH2); 4.36 (q, 2H, OCH2CH3, J=5.6 Hz); 7.19-7.70 (m, 6H, Ar-H); 11.04 (s,1H, indole-NH); 13C-NMR (100 MHz, CDCl3) δ (ppm)= 16.30, 20.29, 21.90, 26.09, 52.20, 102.24, 114.32, 114.98, 123.05, 123.84, 124.95, 125.69, 126.75, 128.03, 128.82, 129.10, 131.02, 132.81, 137.13, 137.83, 138.42, 152.03, 152.92, 164.51, 167.13. Anal. Calcd For C25H20N3O2Cl3 (500.80): C, 59.95, H, 4.02, N, 8.39; Found: C, 59.97, H, 4.04, N, 8.36.
2-Amino-9-bromo-4-(2,4-dichloro-phenyl)-5,6,7,12-tetrahydro-1,12-diaza-dibenzo[a,e]azulene-3-carboxylic acid ethyl ester (7d): Yield; 73%; M.p. 292ºC -294 ºC; FT-IR (cm-1): 3451 (N-H), 3415, 3316 (NH2), 1699 (C=O), 1618 (C=N); 1H-NMR (400 MHz, DMSO) δ (ppm)= 1.21 (t, 3H, J=4 Hz); 1.62 (p,2H, J= 2.8 Hz); 2.29 (t, 2H, J=3.6 Hz); 2.52 (t, 2H, J=8.4 Hz); 3.96 (s, 2H, NH2); 4.40 (q, 2H, OCH2CH3, J=2 Hz); 7.43- 7.66 (m, 6H, Ar-H); 10.72 (s,1H, indole-NH); 13C-NMR (100 MHz, CDCl3) δ (ppm)= 19.45, 21.86, 24.34, 26.83, 48.36, 103.70, 116.59, 117.87, 120.46, 122.42, 123.14, 124.53, 125.11, 125.89, 128.10, 128.96, 132.95, 136.68, 137.22, 137.81, 138.49, 153.14, 154.36, 165.43, 166.28. Analytical Calculated for C25H20N3O2Cl2Br (545.25): C, 55.06, H, 3.69, N, 7.70; Found: C, 55.09, H, 3.66, N, 7.73.
Biological Study
Antimicrobial activity
All the microbial cultures were subjected to analyze their susceptibility/resistance pattern to test samples by well diffusion method [19], using Mueller Hinton agar medium (Cat. No. M1084, HiMedia, India) for bacteria and Sabrose Dextrose Agar medium for fungus. Sterile medium was dispensed into sterile petri dishes aseptically. Enriched both cultures (24 h for bacteria and 48 h for fungus, incubated) were used as inoculums. Using sterile cotton swab, the test organisms were swabbed over the surface of the agar plate aseptically. In each of these plates, wells (10 mm) were cut out using sterile cork borer. The samples were dissolved in the DMF (Dimethyl formamide) and different concentration (20-80 μg/ml) of the sample was loaded onto the wells. Incubate the plates at 37oC (for 24 h for bacteria and 96 h for fungus) upright position of the plates. Solvent was used as control. After the incubation, the diameters of inhibition zones were observed. The studies were performed in triplicate and the average reading was taken. The inhibition zones were compared with those of reference disc.
DNA Cleavage
The DNA cleavage activity of the cyclohepta[b]indole derivatives (6,7a-d) were monitored by agarose gel electrophoresis method [20] on pUC19 DNA. The cleavage was monitored by 10 μl of compounds in tris–HCl buffer and 0.5, 0.6, 0.7, 0.8, 0.9 and 1 μl of pUC19 DNA. The samples with sufficient buffer were incubated for about 2 h in at 37ºC. After incubation, 1 μl of gel loading dye (8 mg of bromophenol blue, 3 ml of glycerol and 10 ml of distilled water) was added to the reaction mixture and loaded onto a 2% agarose gel containing 1 μl of ethidium bromide. The electrophoresis was carried out for about 2 h at 50 V in tris–acetic acid EDTA (TAE) buffer. The bands were visualized under UV light and photographed.
DNA Binding
Binding affinity of the compound with CT-DNA were carried out in double distilled water with tris(hydroxymethyl)-aminomethane (Tris, 5 mm) and sodium chloride (50 mm). The pH was adjusted to 7.2 using hydrochloric acid. A solution of CT-DNA in the buffer gave a ratio of UV absorbance of about 1.8-1.9 at 260 and 280 nm, indicating that the DNA was sufficiently free of protein. The DNA concentration per nucleotide was determined by absorption spectroscopy using the molar extinction coefficient value of 6600 dm3 mol-1 cm-1 at 260 nm. The compound was dissolved in a mixed solvent of 5% DMSO and 95% tris-HCl buffer for all the experiments. Stock solutions were stored at 4°C and used within 4 days. Adsorption titration experiments were performed with a fixed concentration of the compound (10 μm) with varying concentration of DNA (0-50 μm). While, measuring the absorption spectra an equal amount of DNA was added to all the test solutions and reference solution to eliminate the absorbance of DNA itself.
Results and Discussion
Chemistry
The synthetic pathway of the designed compounds 6a-d and 7a-d were shown in Scheme 1. The study was commence by scrutinizing the one pot Knoevenagel condensation reaction of 7,8,9,10-tetrahydro-5H-cyclohepta[b]indol-6-one (1a-d), 2,4-dichlorobenaldehye (2) with malononitrile (3) and ammonium acetate (5) using organocatalyst L-proline yielded corresponding compounds 2-amino-4-(2,4-dichloro-phenyl)-5,6,7,12- tetrahydro-1,12-diaza-dibenzo[a,e] azulene-3-carbonitrile (6a-d). The IR spectrum of 6a displayed absorption bands at 3325, 3251 and 2207 cm-1 assigned to NH2 and C≡N stretching vibrations respectively. Its 1H-NMR spectrum exhibits a broad singlet at δ 4.426 and δ 11.45 ppm corroborating the presence of NH2 and indole NH groups. Aromatic protons materialize in the region of δ 7.08-7.56 ppm and the aliphatic protons shows one pentet at δ 2.37 and two triplets at δ 2.88 and δ 3.27 ppm C6 and C7,C5 protons respectively. The total number of protons matched perfectly with its structure.
In continuation of this work, the next part is to synthesis of 2-amino-4-(2,4-dichloro-phenyl)-5,6,7,12-tetrahydro-1,12-diaza-dibenzo[ a,e]azulene-3-carboxylic acid ethyl ester (7a-d) by one pot Knoevenagel condensation reaction of 7,8,9,10-tetrahydro-5H-cyclohepta[ b]indol-6-one (1a-d), 2,4-dichlorobenaldehye (2) with ethylcyano acetate (4) and ammonium acetate (5) using L-proline as an organocatalyst in dry ethanol. The significant stretching vibrations of compounds 7a-d were proven by IR spectra, which revealed the presence of carbonyl, amino and indole NH stretching frequencies in their respective range. The 1H-NMR spectra of compounds 7a, the amino protons resonates at δ 4.67 ppm. The C-3 substituted ethyl group proton appears at δ 1.73 and δ 5.50 ppm. In addition the appearance of 7 aromatic protons emerged in the range of δ 7.28-7.70 ppm. The 13C-NMR spectra exhibit the appropriate carbon skeleton. The 13C-NMR spectrum revealed the presence of 25 carbons. The structures of all compounds (6a-d and 7a-d) were confirmed by elemental and spectral analyses.
Biological Study
Antimicrobial
A series of compounds (6,7a-d) were tested against bacterial strains such as Pseudomonas aeruginosa, Bacillus licheniformis, Shigella dysenteriae, Klebsiella pneuminiae, Escherichia coli and fungal strains is Aspergillus niger and Candida tropicalis. All the synthesized compounds, except 7b were exhibited good activity against Pseudomonas aeruginosa and compounds 6a, 6a, 7c and 7a were showed moderate activity against Bacillus licheniformis. For the inhibition of bacteria Klebsiella pneuminiae, the active compounds are 6b, 6c, 6a, 7d, 7c and 7b. Among all the synthesized compounds 3b was the one with activity against Shigella dysenteriae bacteria. Furthermore, Compounds 6b, 6d, 7b and 7d moderately inhibited the growth of the Escherichia coli. All the prepared derivatives showed good antifungal activity. The growth of the Aspergillus niger were significantly inhibited by the compounds 6a, 6b, 7a and 7b. Compounds 6c, 6d and 7d were exhibited potent activity against Candida tropicalis. From the results, it was clear that the compounds 6c, 6d, 7c and 7d possess heterocyclic unit with halogen atoms responsible for antimicrobial activities. The zones of inhibition (in mm) of synthesized compounds were summarized in Table 1.
| Zone of inhibition (mm) | ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Name of organism | 6a | 6b | 6c | 6d | 7a | 7b | 7c | 7d | ||||||||||||||||||||||||||||||||
| C | 20 | 40 | 60 | 80 | C | 20 | 40 | 60 | 80 | C | 20 | 40 | 60 | 80 | C | 20 | 40 | 60 | 80 | C | 20 | 40 | 60 | 80 | C | 20 | 40 | 60 | 80 | C | 20 | 40 | 60 | 80 | C | 20 | 40 | 60 | 80 | |
| P.aeruginosaa | - | - | - | 17 | 19 | - | 11 | 12 | 14 | 16 | - | - | 12 | 13 | 15 | - | - | - | - | 16 | - | - | 14 | 18 | 25 | - | - | - | - | - | - | - | 14 | 15 | 16 | - | - | - | - | 17 |
| B.licheniformisa | - | - | - | - | 13 | - | - | - | - | - | - | - | 14 | 16 | 18 | - | - | - | - | - | - | - | - | 15 | 19 | - | - | - | - | - | - | - | 14 | 16 | 18 | - | - | - | - | - |
| K.pneuminiaea | - | - | - | 12 | 18 | - | - | - | 15 | 18 | - | - | - | 13 | 15 | - | - | - | - | 13 | - | - | 13 | 18 | 20 | - | - | - | - | 12 | - | - | 12 | 16 | 19 | - | 10 | 11 | 13 | 15 |
| S. dysenteriaea | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 16 | 22 | - | - | - | - | - | - | - | - | - | - |
| E.Colia | - | - | - | - | - | - | - | 12 | 14 | 16 | - | - | - | - | - | - | 12 | 14 | 18 | 24 | - | - | - | - | - | - | - | 12 | 14 | 16 | - | - | - | - | - | - | - | - | 12 | 14 |
| A.Nigerb | - | - | - | - | 13 | - | - | - | - | 14 | - | - | - | - | - | - | - | - | - | - | - | - | 16 | 17 | 19 | - | 16 | 17 | 20 | 24 | - | - | - | - | - | - | - | - | - | - |
| C. tropicalisb | - | - | - | - | - | - | - | - | - | - | - | - | - | 16 | 22 | - | - | 15 | 16 | 19 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 14 | 16 |
aBacterial strain; bFungal strain
Table 1: Antimicrobial activity of prepared compounds 6a-d and 7a-d
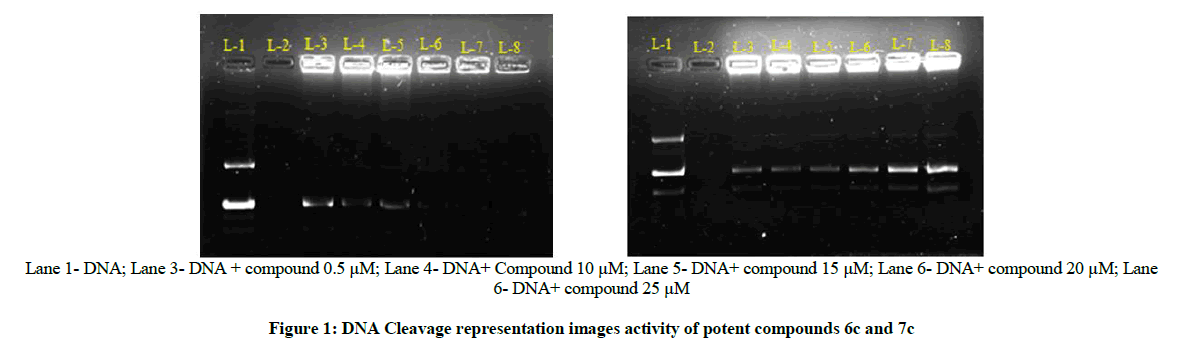
DNA Cleavage
The DNA cleavage activity was determined using gel electrophoresis method. The pictures of the gels are presented in Figure 1. The gel after electrophoresis clearly revealed that, all the tested compounds have been act on the pUC19 DNA as little tailing in the bands can be observed in treated compounds. The difference was observed in bands of all the compounds compared to the control DNA, this shows that the control DNA alone does not show any apparent cleavage as the compounds have been done. It can be concluded that the compound inhibits the growth of the pathogenic organism by cleaving the genome.
DNA Binding
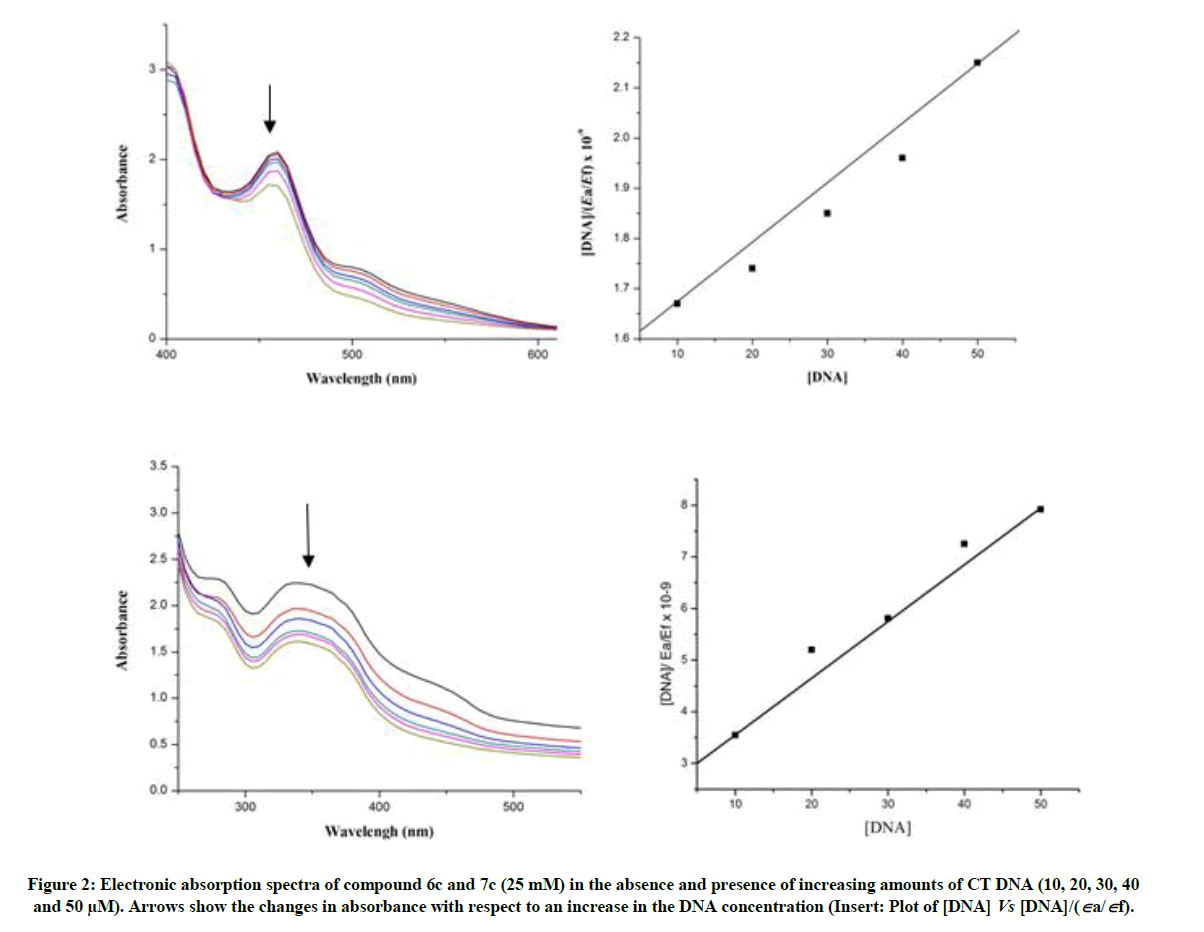
Electronic absorption spectroscopy is one of the most common and effective technique for investigation of the binding modes of organic compounds with CT-DNA. Compound binding though intercalation usually results in hypochromism with or without small red or blue shift, since the intercalative mode involves a strong interaction between the aromatic chromophore and the base pairs of DNA. The binding mode of the compound 6c and 7c has been characterized though absorbance and shifts in the wavelength as a function of added concentration of DNA. Upon addition of increasing amounts of CT-DNA, a significant hypochromism is observed (Figure 2). This can be attributed to a strong interaction between DNA. In order to illustrate quantitatively the consequence, the absorption data were analyzed to evaluate the intrinsic binding constant (Kb), which can be determined from the following equation,
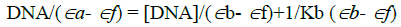
Where [DNA] is the concentration of DNA in base pairs, the apparent absorption coefficient Εa, Εf and Εb corresponds to Aobs/[compound], the extinction coefficient of the free compound and the extinction coefficient of the compound when fully bound to DNA, respectively. From the plot of DNA/(Εa- Εf) versus [DNA], Kb is calculated by the ratio of slope to the intercept. The magnitude of intrinsic binding constant (Kb) value for 6c is 3.74 × 104 and 7c is 6.73 × 104. From the above DNA binding results, it is obvious that the synthesized compound has planarity and extended π system which lead to the possibility of DNA intercalation.
Conclusion
We have described one pot multicomponent reaction method for the preparation of cyclohepta[b]indole derivatives from simple available starting materials. The cyclohepta[b]indole derivatives have been characterized by spectral and elemental analysis. The antimicrobial results revealed that compounds with electron withdrawing groups showed potent activity and found that compounds 6c, 7c, 7d has great inhibitory activity than the others. All of the obtained derivatives showed significant ability to cleave the pUC19 DNA, especially compounds 6c and 7c. These compounds further tested for their DNA binding capability, it showed that the compound has significant hypochomism. This can be attributed to a strong interaction between DNA. These results are suggesting that the synthesized active compounds can be better candidates for future investigations to produce new drugs.
Acknowledgement
The authors are gratefully acknowledged Sophisticated Analytical Instrument facility (SAIF), Kochi and Karunya University, Coimbatore for their technical support in NMR and biological studies. One of the authors A.A thanks the UGC-Hyderabad (MRP No. 4997/14 (SERO-UGC) for financial support.
References
- B. Huang, D. Hu, J. Wang, J.P. Wan, Y. Liu, Tetrahedron. Lett., 2015, 56, 20, 2551-2554.
- M. Beyrati, M. Forutan, A. Hasaninejad, E. Rakovský, S. Babaei, A. Maryamabadi, G. Mohebbi, Tetrahedron., 2017, 73, 5144-5152.
- S.B. Dongare, H.V. Chavan, P.S. Bhale, Y.B. Mule, A.S. Kotmale, B.P. Bandgara, Chin. Chem. Lett., 2016, 27, 99-103.
- Y.H. He, J.F. Cao, R. Li, Y. Xiang, D.C. Yang, Z. Guan, Tetrahedron., 2015, 71, 49, 9299-9306.
- B. Banerjee, Ultrason. Sonochem., 2017, 35, 15-35.
- R. Jiang, Y. Wang, Z. Zhou, Tetrahedron., 2016, 72, 6444-6449.
- G.M. Ariza, N. McConnell, C. Hulme, Org Lett., 2016, 18, 8, 1864-1867.
- C. Pasko, A.A. Dissanayake, B.S. Billow, A.L. Odom, Tetrahedron., 2016, 72, 9, 1168-1176.
- M. Bakherad, A. Keivanloo, S. Jajarmi, Tetrahedron., 2012, 68, 2107-2112.
- B. Li, S. Guo, J. Zhang, X. Zhang, X. Fan, J. Org. Chem., 2015, 80, 11, 5444-5456.
- N.V. Lakshmi, P. Thirumurugan, K.M. Noorulla, P.T. Perumal, Bioorg. Med. Chem. Lett., 2010, 20, 5054-5061.
- S. Gupta, P. Maurya, A. Upadhyay, P. Kushwaha, S. Krishna, M.I. Siddiqi, K.V. Sashidhara, D. Banerjee, Eur. J. Med. Chem., 2018, 143, 1981-1996.
- H. Mirzaei, M. Shokrzadeh, M. Modanloo, A. Ziar, G.H. Riazi, S. Emami, Bioorg. Chem., 2018, 75, 86-98.
- Z. Shi, Z. Zhao, M. Huang, X. Fu, C. R. Chim., 2015, 18, 12, 1320-1327.
- G. Mei, H. Yuan, Y. Gu, W. Chen, L.W. Chung, C.C. Li, Angew. Chem., 2014, 126, 11231-11235.
- P. Goswami, A.J. Borah, P. Phukan, J. Org. Chem., 2015, 80, 1, 438-446.
- E. Yamuna, R.A. Kumar, M. Zeller, K.J.R. Prasad, Eur. J. Med. Chem., 2012, 47, 228-238.
- E. Stempel, T. Gaich, Acc. Chem. Res., 2016, 49, 2390-2402.
- A.W. Bauer, W.M. Kirby, J.C. Sherris, M. Turck, Am. J. Clin. Pathol., 1966, 45, 4, 493-496.
- J. Sambrook, E.F. Fritsch, T. Maniatis, Molecular Cloning, a Laboratory Manual, second ed. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, 1989.